From a humble point of view, as I was attending a bioinformatics and genomics workshop held in FOPCU, the lecturer was pointing to us, that up until now, no one has managed to come up with a method capable of converting a full-functioning protein back into the original nucleotide sequence on its corresponding gene. At that instance, the following thought occurred to me, as to why this would ever be needed?
For starters, we already have the protein in hand, its 3D structure is, for many, completely figured out and some even their orientation in space, their actions and functions. Then, as far as I understand, being the mould from which a protein is later assembled is the only function a gene, or one which is expressed anyways, has. Knowing that for instance, in gastrin hormone, the 4th amino acid is leucine, would it matter whether it was translated from the codon CUA and not UUG?
Now three thoughts impose themselves. I could only imagine that the presence of SNPs (which is basically a nucleotide that varies among individuals and thought to influence certain traits) within the nucleotide sequence of the gene is the reason behind the researchers’ attention. However, this ultimately means, that if a method were to exist, it would have to produce a different nucleotide sequence for proteins coming from different people. Simple logic.
Another probable explanation, that could come to mind, would be the existence of a difference in the structure of the leucine amino acid, held on tRNA molecules with varying anticodons, where each would have some “characteristic” features that distinguish it from the other tRNA. If that were the case, then it probably has managed to fly below the radar for quite some time, as no matter which reference I turn to, it is taken for granted that these amino acids are carbon copies. So being non-identical in any way, would cause the resulting protein to function in a slightly different manner, which could explain the diversity of their actions in varying individuals. Who knows?
Last, but not least, is the possibility of gaining fast insight into the genome of a previously undiscovered species of living organism, where one can quickly figure out all the expressed genes through this simple task of “reverse translation”. However sequences of the unexpressed genes would still have to adopt the old-fashioned way. No choice there!
Just wondering what the future has in store.
Tags: Bioinformatics, fopcu, gene sequence, genome, genomics, nucleotide, protein, research, reverse translation, sequence, translation, workshop
 No Comments »
No Comments »
Hansenula polymorpha, also known as Pichia angusta, with its metabolism highly dependent on methanol as a carbon source, has been excessively employed in the production of therapeutic proteins for the last two decades. The yeast was first discovered in 1950s in spoiled orange juice.

Image source:
http://www.drugdevelopment-technology.com/contractors/contract_research/artes/
The biotechnological interest in Hansenula is mainly attributed to its unique capabilities underlied by its rare characteristics. For instance, being one of the limited group of yeast that are able to assimilate methanol, gives it the advantage of being able to utilize relatively cheap substrates. In addition, Upon high temperature, Hansenula polymorpha shifts its biochemical methanol metabolism pathway to the biosynthesis of trehalose which is a thermo-protective sugar, this fact explains its unique ability to resist temperatures up to 49 degree Celsius. further, Hansenula is able to secrete the protein products directly to the culture, a fact that renders the whole process of downstream processing easier and less costly. finally, the ability to survive in wide pH range,from 2.5 to 6.5, makes it a versatile protein factory which is exploited in the production of various types proteins, each of which requires a very different optimum pH value throughout the fermentation process.

Image source:
http://en.wikipedia.org/wiki/Hansenula_polymorpha
( Budding Hansenula cells)
However, with Hansenula polymorpha post-translational modification processes are not highly regulated, this makes Hansenula useful for the production of relatively small to medium sized polypeptides as Parathyroid hormone, Staphylokinase, Elafin……etc. However, Regarding large polypeptides, mammalian cells with tightly regulated post-translational modification processes represent a better option.
On the industrial scale, different strains of Hansenula polymorpha are being exploited as expression systems. For instance, a genetically engineered strain known as super-transformed strain bearing additional two advantages than the wild type strain. Being super-transformed means that it is capable of secreting Calnexin which is a protein Chaperone that functions to ensure proper folding of the secreted protein. Additionally, Calnexin enhances the protein secretion efficiency of Hansenula . Accordingly, the super-transformed strain gains its industrial potential in terms of quality assurance of the secreted protein product as well as increasing the cost effectiveness of the industrial process.
References: 1.“Hansenula polymorpha” wikipedia, www.wikipedia.org
2. Gellissen G, “Hansenula ploymorpha : Biology & Applications”, 2002
3.Marcos A. Oliveira, Victor Genu, Anita P.T. Salmazo, Dirce M. Carraro and Gonçalo A.G. Pereira1 ”The transcription factor Snf1p is involved in a Tup1p-independent manner in the glucose regulation of the major methanol metabolism genes of HansenulaPolymorpha”, Genetics and Molecular Biology, 521-528 (2003)
4. Giuseppinia Parpinello, Enrico Berardi, and Rosanna Strabbioli,” A Regulatory Mutant of Hansenula polymorpha Exhibiting Methanol Utilization Metabolism and Peroxisome Proliferation in Glucose”, J Bacteriol. 1998 June; 2958–2967.
Tags: Calnexin, Chaperone, Downstream processing, Elafin, Hansenula polymorpha, methanol, Parathyroid hormone, Post-translational modification, Staphylokinase, Super-transformed, Therapeutic proteins, thermo-protective, trehalose, Yeast
 No Comments »
No Comments »
Dr. Betsey Dexter Dyer is a professor of Biology at Wheaton College, Massachusetts, USA. She received her Ph.D.  from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including
The Modern Scholar: Unseen Diversity: The World of Bacteria (Audiobook), The Origin of Eukaryotic Cells (Benchmark Papers in Systematic and Evolutionary Biology, 9) (1986), Tracing the History of Eukaryotic Cells (1994 – with Robert A. Obar), Explore the World Using Protozoa (1997 – coauthor), A Field Guide to Bacteria (2003) and Perl for Exploring DNA (2007 – with Mark D. LeBlanc).
I recently read about her book “A Field Guide to Bacteria” and wanted to know more about her view on “Bacteriocentrism,” and how one becomes “bacteriocentric.” I was so lucky to be introduced to Dr. Dyer, who was kind enough to accept to be interviewed for Micro Writers, and immediately answered my questions about her book and her student project, Genomics Research Group, via email. So, here I am, sharing with you this very interesting interview.
1. Why did you decide to write the book “A Field Guide to Bacteria”? Field microbiology and symbiosis are among your research interests ; is this one of the reasons that made you dig deeper in bacterial populations and “think like a microbe”?
I first got the idea of a field guide when I was a graduate student and was very fortunate to be on a field microbiology expedition (in Baja California Mexico) with some world famous field microbiologists. I realized at once that a field guide should be written and one of them should write it and probably would! It did not occur to me that I would write it. It took me years to get to the point of being secure enough with my career. First I had to get a PhD, then a job, and then tenure. I also got married and had two children. Finally, about 15 years after that original idea, I realized that I had been accumulating enough information that I should begin to write. And so I did. However, I am still a bit surprised that nobody else wrote it.
I am naturally drawn to tiny things. I got a microscope for a present when I was 11 years old and it transformed some of my views of biology. I found that I loved the microscopic world. But I also like miniatures in general such as tiny furniture and dishes and things in dolls houses. I have in my library at home, some shelves devoted to a doll house and two miniature rooms.
2. What is meant by “becoming bacteriocentric”? And how does this lead to better understanding of the biology of bacteria?
We humans are mostly visual and auditory, the primary senses by which we perceive and analyze the world. It is probably impossible for us to be otherwise. Furthermore, we are gigantic and multicellular and terrestrial in marked contrast to the vast majority of organsims on Earth. Nonetheless, I think it is an excellent exercise for any biologist at least to try bacteriocentricity. The bacterial or microbial world is primarily olfactory and tactile. They are single celled (intimate with their environments), tiny and aquatic. I cannot avoid being anthropocentric but I can at least be more aware of the limitations of my size, habitat, and senses. My goal is to have as much humility as I can manage when I observe the world of microbes.
3. “Many groups of bacteria can be easily identified in the field (or in the refrigerator) without a microscope” and “Bacteria can be seen and smelled”, as a pharmacy student, I want ask how could that be achieved?
Well, do you have the book yet? There are many examples but the basis of all of them is that bacteria, when they are in an appropriate environment, are likely to do quite well: reproducing abundantly, taking in and transforming molecules, sending out wastes. In many cases (surprisingly many) the abundance is on a level perceptible by humans. The field marks just need to be revealed and interpreted. Otherwise, they may be easily overlooked or misunderstood. My first experience with this as a graduate student was being shown the distinctive pigmentations and odors of bacteria in sulfur cycles in Baja California.
4. You have a project with Dr. Mark LeBlanc, professor of Computer Science, called “Wheaton College Genomics Group.” Why did you do such a project for undergraduate students? How did it raise their potential?
One day about ten years ago, Mark asked me if I had any large datasets that his computer science students might analyze in their course on algorithms. It happens that I teach genetics and am fascinated with genomes. At that time, genome sequences were becoming more available at NCBI. I had not realized that it would be so easy to collaborate with a computer scientist. I had lots of questions about genomes and we just started right in with devising some answers. Right now, we are interested in characterizing horizontal transfer events of distantly related bacteria and archaea. There are hundreds of complete microbial genomes at NCBI and most have not been completely analyzed. Therefore, there is plenty for us and our students to do.
We ended up writing a book on the topic because we wee in need of a text that could be used both by biologists and computer scientists.
Tags: A Field Guide to Bacteria, bacteriocentric, bacteriocentrism, Betsey D. Dyer, biologists, computer scientists, field microbiology, Mark LeBlanc, NCBI, Perl, Perl for Exploring DNA, sulfur cycles, symbiosis, Wheaton College Genomics Group
 1 Comment »
1 Comment »
Yediot Ahronot (Literally: Latest News)
Cure for radiation sickness found?
Published: 07.17.09
A team of scientists has succeeded in developing an anti-radiation. “The process that led up to the medical innovation dates back to 2003, when Professor Gudkov –the head of the team- came up with the idea of using protein produced in bacteria found in the intestine to protect cells from radiation.” Mice received that purified protein survived the amount of radiation that killed the control group.
What kind of news is that?! Do that legendary bacteria and that miraculous protein actually have names?! If so, why don’t newspapers include it? Should bloggers do everything?!
Professor Andrei Gudkov – Chief Scientific Officer at Cleveland BioLabs, is interested in protecting cells against apoptosis induced by cancer therapy as well as radiotherapy. He worked on p53 –the famous tumor suppressor- and found that p53 has a role in inducing apoptosis. The research group suggested that p53 inhibitors can protect normal cells against chemo- and radiotherapy, and it’s been found that it sensitizes tumor cells to the therapy (PMID: 15865929). They also showed that PFTmu (pifithrin-mu), a small isolated inhibitor of p53, protected primary mouse thymocytes from p53-induced apoptosis caused by radiation (PMID: 18403709).
The breakthrough discovery mentioned above has been published in Science – 11 April 2008. It is about the injection of “flagellin” purified from Salmonella enterica serovar Dublin into mice and monkeys. It causes suppression of apoptosis by binding to Toll-like receptor 5 (TLR5) and activation of the nuclear factor–kappaB (NF-kappaB) pathway, the same mechanism used by tumor cells to inhibit the function of the p53 pathway (PMID: 18403709). To reduce its immunogenecity and toxicity, they engineered a polypeptide derived from flagellin with the “important” domains only, N and C termini separated by a linker. The engineered protein (named: CBLB502) was found to provide radioprotection in rhesus monkeys and mice against lethal doses of gamma-radiation and accompanied hematopoietic system and gastrointestinal tract acute radiation syndrome, with no alteration of the efficacy of the radiotherapy.
Tags: Andrei Gudkov, anti-radiation, flagellin, NF-{kappa}B, p53 inhibitor, PFTmu, pifithrin, radioprotective, radiotherapy, Salmonella enterica, TLR5, Yediot Ahronot
 No Comments »
No Comments »
Not too long ago, I read about a research done at the Kennedy Institute of Rheumatology Division, which has identified a new ligand for Toll-like receptor 4. This receptor was previously known for activating the immune system through the detection of threats as lipopolysaccharide or gram-negative bacteria. The new ligand, Tenascin-C, is an extracellular glycoprotein, whose elevated expression in cases of inflammation provoked scientists to study its role in the process. 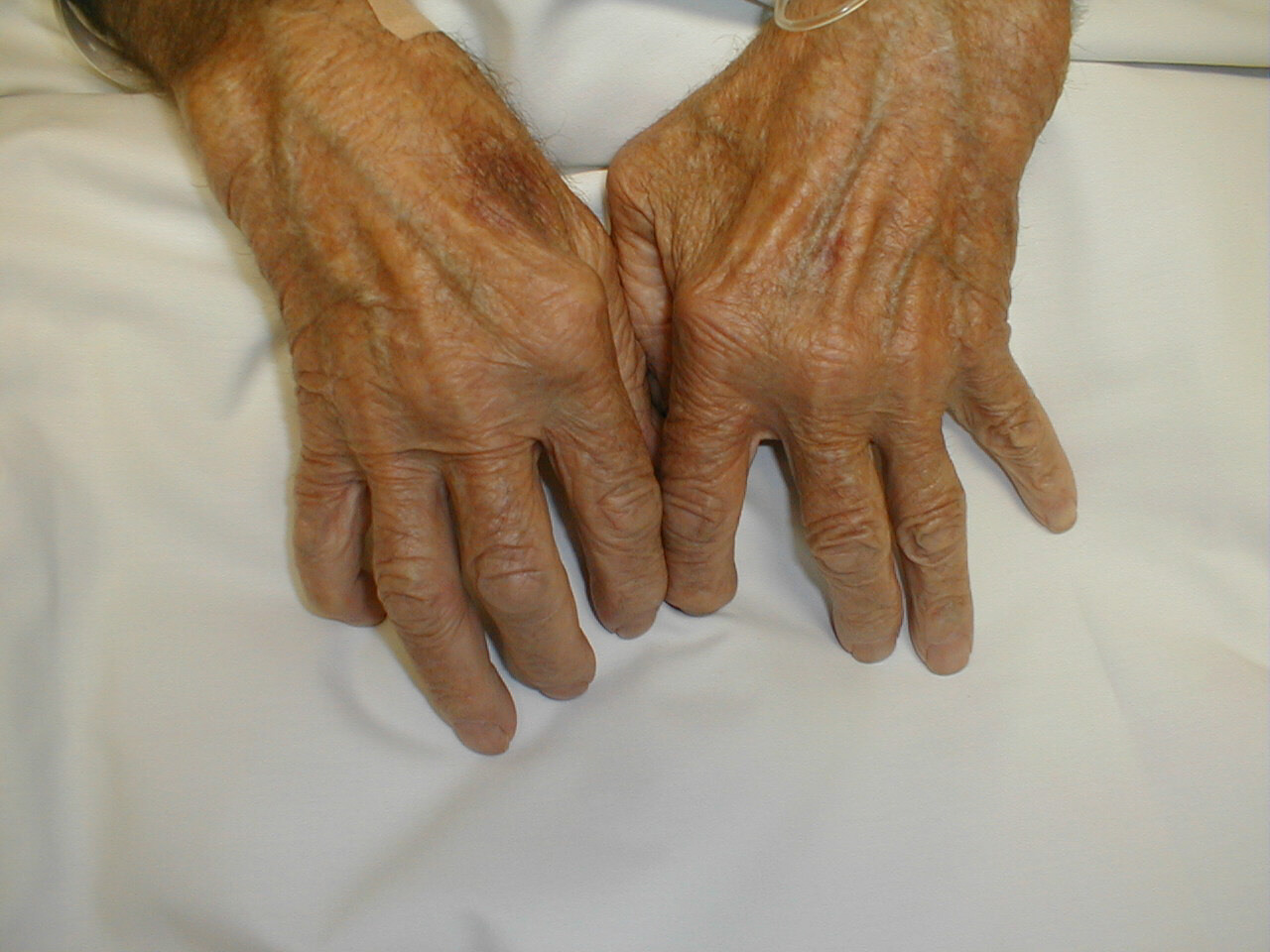
The study noted that its presence was critical to maintain the ongoing inflammation seen in cases of rheumatoid arthritis. In reference to this study, the author stated “We have uncovered one way that the immune system may be triggered to attack the joints in patients with rheumatoid arthritis. We hope our new findings can be used to develop new therapies that interfere with tenascin-C activation of the immune system and that these will reduce the painful inflammation that is a hallmark of this condition”
I was able to contact Dr Kim Midwood and obtained this brief interview:
1. Do you have any speculations as to why Tenascin-C is overly expressed in certain individuals causing prolonged inflammation cases, whilst remaining within normal levels in others?
What regulates tissue levels of tenascin-C is not currently known and this is something that we are working on finding out.
2. From the different ligands of TLR4, why was Tenascin-C of particular interest in your research?
I have a long standing interest in how cell behavior is influenced by the extracellular environment, and in particular the role of extracellular matrix proteins in regulating cell phenotype during the response to tissue injury. For the last 10 years, I’ve been studying the role of tenascin-C – a protein specifically and transiently expressed upon tissue injury, but persistently expressed in chronic inflammatory diseases such as rheumatoid arthritis. This pattern of expression, plus the high homology of tenascin-C domains to other known pro-inflammatory matrix molecules or ‘DAMPs’ prompted us to investigate whether tenascin-C was an endogenous activator of the immune response and whether its persistent expression in RA contributed to disease pathogenesis.
3. What do you think the extent of similarity will be between the mice & human response to the Tenascin-C blockage?
I cannot predict how differently the mouse and human will behave.
4. Do you suspect a certain mechanism of the increase in inflammatory molecules caused by Tenascin-C?
We know that tenascin-C activates TLR4, activation of this receptor is well known to induced the expression of pro-inflammatory genes via activation of many intracellular signaling pathways.
5. How do you see the potential of such study for rheumatoid arthritis patients?
We plan to identify ways to inhibit the pro-inflammatory action of tenascin-C in the hope that this may be useful in reducing chronic inflammation in the joint.
Original research paper: Tenascin-C is an endogenous activator of Toll-like receptor 4 that is essential for maintaining inflammation in arthritic joint disease. Nature Medicine 15, 774 – 780 (2009). PMID: 19561617 (Vote for the abstract on Biowizard)
Image Credit: Davidson College Undergraduate Course
Tags: chronic inflammation, DAMPs, endogenous, inflammation, Interview, intracellular signal, Kim Midwood, ligand, pro-inflammatory, rheumatoid arthritis, rheumatoid therapy, Tenascin, TLR4, Toll-like receptor
 No Comments »
No Comments »
A team of Harvard scientists has taken the first step to solve mystery about why HIV patients are more susceptible to TB infection.
According to USAID, 42 million people are HIV infected & almost one third of them are also TB infected. It is believed that HIV interferes with the cellular and molecular mechanisms used by the lungs to fight TB infection. 
This mystery has been solved when scientists extracted immune cells called “alveolar macrophages” from the lungs of asymptomatic HIV +ve patients as well as healthy patients ” HIV –ve.” They observed a decrease in response towards TB bacterium in HIV +ve patients when compared to HIV –ve patients.
A further examination of lung specimens showed an increased level of a molecule called IL-10, which elevates the amount of a protein called “BCL-3” in alveolar macrophages and this reduces their ability to ward off TB infection.
It seems that HIV increases severity of TB infection, where both represent two of the most significant health challenges in human history.
Source: Science Daily.
Image credits: The HIV replication cycle.
Tags: BCL-3, HIV, IL-10, TB, USAID
 No Comments »
No Comments »
When you hear/ read the term “Phage Therapy“, you’ll be automatically directed to the concept of using bacteriophages, the virus-like particles that infect bacteria, to kill/ lyse the resistant bacterial strains, instead of the “useless” antibiotics that allowed bacteria to fool them & develop resistance against them. The initial target of phage therapy was to kill the bacteria using phages; because they act like any other virus; get in, multiply and lyse the cell. But, by this way, bacteria develop resistance against phages more rapidly. So, they may become useless by time. In this paper from PNAS: “Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy,” two bioengineers, Timothy K. Lua and James J. Collins, from Boston University successfully engineered the Enterobacteria filamentous phage M13 to weaken bacteria not to kill it. Sounds strange, right? By engineering M13, they gave us a variety of options:
1st, we may make M13 overexpress a bacterial protein named lexA3 which inhibits the ability of the bacteria to repair their damaged DNA by the action of Ofloxacin –as pharmacophils, who had 2 consecutive chemotherapeutics courses, we may recall that quinolones’ MOA is generation of ROS. So, the repressor suppresses the bacterial SOS mechanism. Very promising results were observed; the adjuvant therapy increased the survival rate of mice infected with resistant E. coli. It was also observed that the adjuvant therapy reduced the rate of developing mutations/ resistance within the E. coli population.

2nd, bacteriophage can be responsible for expression of certain proteins that can attack gene networks in bacteria which are not target for existing antibiotic classes. I will mention just one example here, expression of CsrA which is a “global regulator of glycogen synthesis and catabolism, gluconeogenesis, and glycolysis, and it also represses biofilm formation,” biofilms is thought to be related to antibiotic-resistance and OmpF porin which is used by quinolones to enter the bacterial cell, it may enhance its entrance.

Now, thanks to the engineered phages, we can use the old beloved antibiotic classes to treat bacterial infection using the engineered phages as an adjuvent therapy to potentiate the cidal action of the antibiotic on the former-resistant strains. A precaution was made to ensure that no lysogeny would take place in the human cells is that the phages were engineered to be “nonreplicative”. But we still have two problems regarding Phage Therapy in general: identifying the strain responsible for the infection & making sure that the human immune system won’t elicit an immune response against phages, they’re “foreigners” after all!
Image credits:
1- “Schematic of combination therapy with engineered phage and antibiotics. Bactericidal antibiotics induce DNA damage via hydroxyl radicals, leading to induction of the SOS response. SOS induction results in DNA repair and can lead to survival. Engineered phage carrying the lexA3 gene (lexA3) under the control of the synthetic promoter PLtetO and an RBS acts as an antibiotic adjuvant by suppressing the SOS response and increasing cell death”: http://www.pnas.org/content/106/12/4629.figures-only
2- “CsrA suppresses the biofilm state in which bacterial cells tend to be more resistant to antibiotics. OmpF is a porin used by quinolones to enter bacterial cells. Engineered phage producing both CsrA and OmpF simultaneously (csrA-ompF) enhances antibiotic penetration via OmpF and represses biofilm formation and antibiotic tolerance via CsrA to produce an improved dual-targeting adjuvant for ofloxacin”: http://www.pnas.org/content/106/12/4629.figures-only
Tags: antibiotic, bacteriophage, bioengineering, engineered bacteriophage, M13, ofloxacin, phage, phage therapy, resistant
 2 Comments »
2 Comments »
This is not a prison break scheme. I was shocked to hear on BBC that researchers published a study, in the  journal Genetic Vaccines and Therapy, about a new route for the delivery of specifically DNA virus vaccinations. Using the vibrating needle, normally used in tattoo parlors, they first experimented with mice & found a 16-fold increase in the humoral & cell-mediated antibody response elicited by these animals compared to the intramuscular injection. The needle implants small DNA fragments into the epidermis, which triggers a non-specific immune response believed to be the reason for the higher antibody levels found despite the lower dose of DNA used. journal Genetic Vaccines and Therapy, about a new route for the delivery of specifically DNA virus vaccinations. Using the vibrating needle, normally used in tattoo parlors, they first experimented with mice & found a 16-fold increase in the humoral & cell-mediated antibody response elicited by these animals compared to the intramuscular injection. The needle implants small DNA fragments into the epidermis, which triggers a non-specific immune response believed to be the reason for the higher antibody levels found despite the lower dose of DNA used.
Flattering as it sounds, a lot of skeptics doubt that it would become a complete replacement of the conventional routes currently in use. It does not come without a cheap price either. Many tattoo lovers loathe the accompanying pain. Plus, potential users won’t be getting a tattoo in the process either because the needle won’t be loaded with ink.
Just truly amazed at whoever first comes up with such ideas and tries putting them to the test.
Image Credit: Enquirer
Tags: DNA, Gene Therapy, immune response, tattoo, vaccine
 2 Comments »
2 Comments »
Meeting a scientist is like reading many books in few minutes, books of science and books of life. To me, though, meeting Dr. Niyaz Ahmed (1) was like reading a library. He is one of the most active advocates for Open Access (OA) and open evaluation of science. His support for OA is not just by words, but through sharing effectively in such purpose as he is a section editor of Microbiology and Genomics in PLoS ONE and a chief editor in Gut pathogen, the official journal of ISOGEM (the International Society for Genomic and Evolutionary Microbiology), in Sassari, Italy, of which Dr. Ahmed is a co-founder and the General Secretary. ISOGEM members work on developing post-genomic ideas to serve the public health and the environment. Dr. Ahmed is one of the faculty members of Faculty of 1000 Biology, the expert guide to the most important advances in biology.
Dr. N. Ahmed’s early start was in India as he graduated in Veterinary Medicine and obtained his Masters degree in Animal Biotechnology, then his PhD in Molecular Medicine. Currently, he is also based in India, in Hyderabad University as an Associate Professor of Biotechnology and a staff scientist in the Center of DNA Fingerprinting and Diagnostics in Hyderabad. This is not everything about him, his homepage reveals more, especially those pioneer achievements in the research world in India, like being a co-principal investigator of the Mycobacterium W genome program, India’s first whole genome sequencing project, and the most amazing part of his research is working on Helicobacter pylori chronological evolution and phylogeographic analysis.
Having read about all these achievements, I had many questions in mind, but, knowing how busy he is, I tried to reduce them to the following few questions that cover different areas of his activities. Dr. Ahmed generously agreed to answer them all.
Dr. Niyaz Ahmed, any microbiologist would be honored to meet you, and so am I. I am one of the “Micro Writers” blog members; we are young enthusiastic students from Faculty of Pharmacy, Cairo University in Egypt, mostly undergraduate and some graduate students, who are looking from the blog window towards the research world. Most of us wished someday to become scientists; “Micro Writers” is like the first real step on the right way. Our blog focuses on different areas of microbiology. It is like a message by students so far from Egypt, but hopefully from other places as well, to students all over the world. I read a lot about your achievements, especially those concerning the H. pylori phylogeographic analysis. As you have a great experience in research, reviewing articles and especially because you own a blog, I would like to ask you a few questions, and get them published with your answers on the blog, in order to teach and guide not only us, but the all students from everywhere.
Since you are working on the population structure of H. pylori, and on their chronological evolution, can the phylogeographic analysis of bacterial strains lead to infection control or innovative treatment? If yes, how?
-
N.A: It may not directly lead to product development as such, but, it can definitely explain how the spread-patterns of this organism are? and that has a lot of bearing on infection control in terms of tracking the infection sources. Also, knowing separate lineages in greater details offers accurate diagnostic development which in turn helps keep a tab on their emergence or otherwise.
If phylogeographic analysis can teach us about migration of some strains from one place to another, can we use it in a reverse way? I mean can we use this analysis to learn about the origin of mankind, and his migration routes?
-
N.A.: Indeed, we can do that. For a bacterial parasite to become a surrogate marker of human history (including migrations), it needs that its population structure mimics with that of the host. It is now established that H. pylori has most probably coevolved with its human host and thus its population structure is juxtapose to the human population structure. Therefore, we can use it as a marker of human history and geography.
Concerning the unpublished ideas of your studies, which suggest that bacterial proteins may be able to perform both the immune stimulatory and immune evasion tasks, the simple question is, do you think bacteria are smart? Do they have that smart controlling machinery that is able to distinguish between all these factors?
-
N.A.: The term ‘smart’ is reserved for humans. However, we can say that bacteria are highly adaptable. Several bacterial traits such as quorum sensing, community interactions (such as that seen in biofilm formation), molecular mimicry, immune evasion, dormancy etc. definitely point towards their being ‘intelligent’.
You talked about the host-pathogen relationship. I would like to add one more suggestion, which is that bacteria may face the immune system or the host body as a community of workers, not as independent cells. It is like the idea of sociomicrobiology in biofilms. Does this social behavior apply to other communities of bacteria other than bioflims, e.g. the community of H. pylori cells against the human defense system in the stomach?
-
N.A.: Indeed there is a body of evidence coming in via both in vitro and in vivo observations that H. pylori cells tend to form biofilms which is favored by gastric mucin and by mutations of luxS, and the cagE type IV secretion gene. This observation is important to understand H. pylori’s resistance to host immune responses and antibiotics, and in microenvironmental pH homeostasis which conditions and streamlines the growth and survival of H. pylori in vivo.
You mentioned at your homepage that strains of H. pylori in the Southern Asia are not life-threatening in the same time they may cause gastric cancer in the West. Do you think the lifestyle in Southern Asia, especially the food (extra-spicy food), might lead to evolution of these non-lethal strains, and vice versa in case of the western lifestyle?
Adoption of open access (OA) in science, isn’t it a huge responsibility, with great challenges, most probably the funding challenge? I mean, who will pay always for open access science?
-
N.A.: Open Access is a reality and the most practical approach to augment knowledge sharing towards science education, research and the practice of medicine and agriculture. Developing countries that have brains but no library budgets and that have been until now deprived of access to knowledge will be at a great benefit with this OA revolution. I do not see any challenge here as funding agencies such as NIH, Howard-Hughes Institute and Welcome Trust have already started subsidizing costs for OA in a big headway.
You started in India; did you feel some bias (in the international journal publishing) towards some western countries (e.g., USA, Germany, France, UK, etc.)? Do you think that the researchers from the developing countries should be encouraged in a way or another, especially against the fees and restrictions of publishing?
-
N.A.: I firmly believe that originality and novelty have no barriers. If your ideas are novel and your findings well defended, no one can stop them from seeing light of the day. Developing country scientists are equally respected – I can cite my own example, I was made Section Editor at PLoS ONE to oversee an extremely important area, microbial genomics; for this appointment, my affiliation and my geographical co-ordinates were not the important criteria but my standing in the field of medical microbiology, the efficiency with which I handled the issues and my professional commitment to the cause and ethos of OA publishing. However, there may be exceptions of which I have no idea. Nevertheless, this is a free world and if one has zeal and dedication, success is not that far. These days, publication restrictions based on novelty or being on ‘cutting edge’ or otherwise, are no issues. Developing country scientists are thus free to bring forward their research even on topics which may not be interesting to a few ‘glamour magazines’ of science, but they are welcome at descent venues such as PLoS ONE; they are free to even send in aspects of their folklore medicine, their local environmental problems, cropping, local biodiversity, energy production, their cultural and human anthropological aspects (and topics cutting across these) wherein no one is their competitor! Fees for publication is not a problem, PLoS and Biomed Central both waive off publication charges quite generously in almost all cases of genuine inability to pay.
In an old interview, you said that blogs are a good chance for young scientists to express their ideas for free, what about the older scientists in the developing countries who need more space to express their research activities, in the same time so many restrictions are imposed on them to get their papers published in the international journals?
-
N.A.: Old, and young, all need to be part of the dialogue. Even for busy academics, blogs are extremely helpful although they are no alternative to a proper publication of results or viewpoints. Blogs can effectively blend zeal and enthusiasm of ‘young Turks’ with the experience and patient advice of the ‘old guards’. Academic institutions in the developing countries should support such portals wherein students and faculty discuss research to arrive on novel interpretations and provocative ideas in an interdisciplinary environment of learning and enabling.
As an editor and reviewer in so many journals, what is your advice to some young Egyptian researchers like us in the “Micro Writers” blog who work with the fewest capabilities and with minimal resources, to get our papers published in an easier way?
-
N.A.: I love the Micro Writers Blog. You are the torch-bearers for all other undergraduate students in our countries of the developing world. My advice is try hard to deliver your best. Do not write on complaints – write about solutions! Do not present as Egyptian, or African, think globally, be a part of global community in the best interests of humanity, the environment and wellbeing of all creatures. I can quote from the Holy Qur’an: “Wallatheena Jaahadu Feena Lanahdiyannahum subulana” (The more you dedicate yourself and strive hard, the more will be the opportunities and paths widened for you from the Almighty). (2)
Finally, I would like to thank you Dr. Niyaz for the time you gifted for us to answer my questions with plentiful ideas, objects, and words. I am quite sure, each one who reads this interview will get benefited more than once.
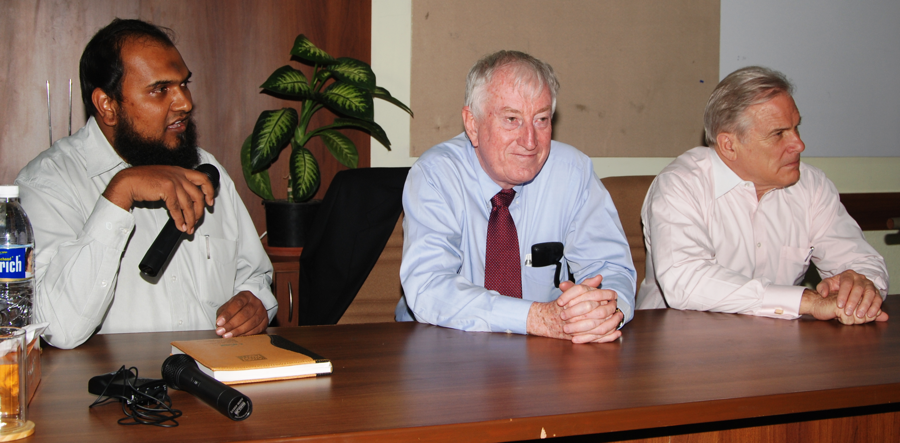

Dr Niyaz Ahmed (left) co-ordinating a ‘student-scientist-administration’ meeting with Nobel Laureate Prof. Peter C. Doherty (center) and Australian High Commissioner to India, Mr John McCarthy (right).
———
(1): Dr Niyaz Ahmed, PhD
General Secretary, ISOGEM
Section Editor, PLoS ONE
Chief Editor, Gut Pathogens
Assoc. Editor, Ann Clin Microb Antimicrobials
Assoc. Editor, Acta Veterinaria Scandinavica
Faculty Member, Faculty of 1000 Biology
Corres. Fellow, Eur Helicobacter Study Group
(2): EL-Qur’an El-Karim, chapter #29 (El-Ankaboot), verse #69.
Image credits: Dr. Niyaz Ahmed.
Tags: Academic institutions in the developing countries, bacterial dormancy, biofilm, Biomed Central, cagE type IV secretion gene, chronological evolution, developing countries, Egyptian researchers, Faculty of 1000 biology, gastric cancer, gastric mucin, Gut pathogen, Helicobacter pylori, host-pathogen relationship, Howard-Hughes Institute, Hyderabad University, immune evasion, ISOGEM, luxS, microenvironmental pH homeostasis, NIH, Niyaz Ahmed, open access, phylogeographic analysis, PLoS ONE, quorum sensing, smart bacteria, the Center of DNA Fingerprinting and Diagnostics, Wellcome trust
 3 Comments »
3 Comments »
Microbiology, Immunology & Biochemistry Dept.*
Faculty of Pharmacy
Cairo University
Bioinformatics Practical Exam – Winter 2010**
Time allowed: Lab computers will automatically hibernate after 2 hours.***
Target: Assigning the function of the uncharacterized protein O67940_ AQUAE from Aquifex aeolicus ****
A suggested procedure:
1- Get the amino acid sequence of the protein from UniProtKB
— Run it through BLAST to find homologs (related sequences). Do not forget to choose Blastp & PSI-BLAST
— Check the assigned hits (known function & solved crystal structure) which have highest possible similarity (highest score/ highest % id) to your query.
2- Check obtained BLAST alignment of those proteins against your query.
3- Check if the protein belongs to any protein family using PIRSF & COGs
— Check if the protein shares any conserved domain with assigned function using Pfam.
— Using PROSITE the functional site database, check if the protein shares any sequence motifs with other proteins
4- Check if the protein belongs to a superfamily using SCOP database, which provides structural and evolutionary relationships between proteins.
5- As you don’t have the crystal structure of your Aquae protein & you have the structure of the closest assigned protein, use VAST to search & align protein related structures to yours.
6- Extract homologs.
7- Multiple alignment (structure-guided alignment) using Cn3D
— Neighbor-joining (NJ) phylogenetic analysis using CDTree
8- Use PDBSum to obtain an overview of the protein–ligand interactions available for your query.
9- Alignment of homologous sequences to identify conserved functional residues.
10- Evidence-based assignment of biological function of query O67940_Aquefix.
🙂GOOD LUCK 🙂
* What have I got to lose?!
** I have faith.
*** I can provide that; I know a guy who knows a guy!
**** Frankly, I wanted to pick a different protein, but I hesitated.
Tags: alignment, Bioinformatics, Cn3D, COG, crystal structure, functional site, homolog, motif, PDBSum, Pfam, phylogenetic analysis, PIRSF, PROSITE, protein family, PSI-BLAST, related structures, SCOP, superfamily, uncharacterized protein, UniProtKB, VAST
 9 Comments »
9 Comments »
|























 Entries (RSS)
Entries (RSS)