Archive for the “Molecular Sciences” Category
Though risky, transplants are crucial treatment interventions for many patients.They can save lives of cancer patients, others with severely ill organs and recently there are trials to make them a mainstream treatment for autoimmune disease patients ( i’ve just read an article about that topic and i’d really love to write about it soon too).
But the major problem with transplants, other than the agonizing wait for the right donor on endless lists for sometimes many years, is that when things go wrong with transplants, the patient’s life becomes at mortal risk. Almost 40% of transplant patients will show rejection episodes within the first year after the operation. The detection of these immunological reactions are usually so late, and the only solution will be to flush the patient’s system with huge doses of immunosuppressive drugs that are toxic themselves and can have debilitating effects on cancer patients for instance. Also, to be able to detect rejection reactions, the doctors should take biopsies of the new organ, a process that can cause damage to the organ itself, let alone the stress and the already fragile patient condition. Transplant patients have to undergo exploratory biopsies monthly for one year after the operation!!!!!
 Will these risky, life-saving procedures be safer in the future? Early detection of these reactions was an interesting topic and a field of research for the cardiologist Hannah Valantine of Stanford University School of Medicine in Palo Alto, California. In 2009, she devised a new test that detects the immunological changes in a transplant patient in an episode of rejection. The test, called AlloMap, became the first of its kind to be approved by the FDA for use in the detection of heart transplant rejections. Yet, it failed to detect the rejections early in about half the patients.This, of course, didn’t satisfy Valantine.
Along with biophysicist Stephen Quake of Stanford, they came out with a much more sensitive test. The idea was that DNA from the new organ constitutes around 1% of the free DNA in blood of transplant patients. This DNA is foreign from the DNA of the patient and using her test, it can be very sensitively detected, despite the fact that it is circulating in minute amounts. To validate their test, they used it on stored plasma samples from transplant patients that later showed rejection signs. It was found that the amounts of the rejected organ’s DNA in such episodes are elevated soon after the surgery and constitutes around 3% of the free DNA in plasma, and of course, will be much elevated later, in the peak of the episode. They reported the results in The Proceedings of National Academy of Sciences.
The good thing about this test, besides its high sensitivity, is that it is much less invasive than a biopsy, and the biopsy will not be needed except for confirmation, in case the test is positive and the DNA % is higher than normal. Also, early detection will allow doctors to use much smaller doses of immunosuppressives to control the case and therefore, less side effects will be experienced. Valantine is a cardiologist, but believes the test can be used with other types of transplanted organs, other than hearts.
Source: ScienceMag
Tags: AlloMap, cardiology, FDA, immunosuppressives, organ rejection, transplants
 No Comments »
No Comments »
 After the Human Genome Project was successfully completed in April 2003 and it was assured that humans are identical in the sequence of their genome by 99.9%, researchers are moving on to find more about the 0.1% left. Although our genome is made out of 3 billion bases (A’s, G’s, C’s and T’s), the 0.1% genetic difference is extremely significant. This is due to the fact that this small percentage holds the key to most of the consequences of such a variation between human beings (e.g susceptibility to diseases and also response to drugs). This introduced the term SNPs (single nucleotide polymorphism) which was found to be highly involved in variation of response to many drugs (like anti-cancers) and the list is increasing every day. After the Human Genome Project was successfully completed in April 2003 and it was assured that humans are identical in the sequence of their genome by 99.9%, researchers are moving on to find more about the 0.1% left. Although our genome is made out of 3 billion bases (A’s, G’s, C’s and T’s), the 0.1% genetic difference is extremely significant. This is due to the fact that this small percentage holds the key to most of the consequences of such a variation between human beings (e.g susceptibility to diseases and also response to drugs). This introduced the term SNPs (single nucleotide polymorphism) which was found to be highly involved in variation of response to many drugs (like anti-cancers) and the list is increasing every day.
To be more focused on the human variations represented by the 0.1 percent, the NIH led the international HapMap project. Their work was performed on four populations groups: the Yoruba people in Ibadan Nigeria,the Japanese in Tokyo, the Han Chinese from Beijing and Utah residents from western and northern Europe. The work began in October 2002 and successfully ended in October 2005. The work of three successive years helped the researchers invent a shortcut for studying SNPs. Scientists believe that there are about 10 million SNPs distributed among 3 billion base pairs which make up our genome, so scanning the whole genome of millions of people for such SNPs would be extremely expensive. After the HapMap project, researchers demonstrated that variants usually tend to cluster into neighborhoods (called haplotypes) and thus the number could be reduced to 300,000 SNPs only. This means that they could reduce the work load by about 30 folds.
The genome-wide association (GWA) studies aim to pinpoint the genetic differences, which cause a certain disease (or a biological trait) by comparing a group of people (who have the trait under research) to a control group (people who are free from this trait). Utilizing thousands of SNPs markers, we can identify regions (loci) which are statistically different between patient and control groups. Thus, we can identify the genetic difference between sick and control people, even though the difference was subtle. This means that the combination of slightly altered genes plus environmental factors could be well studied. The conventional ways, usually used to study genetic differences, are mainly based on selecting the candidate gene based on knowing or suspecting the mechanism of the disease. GWA helps scanning of the whole genome in a comprehensive unbiased manner. It will let us get the whole picture about other, non expected, contributing genes. In this way, GWA studies will help us study the multi-factorial diseases (like cancer and diabetes) in a more rationalised way.
Another challenge has come up: What about the genetic variations due to geographic ancestry? It is also a significant contributing factor to variation among humans and all the efforts are directed towards making a somewhat universal map of human genome to help develop individualized drugs. A group of scientists led by David Reich, an assistant professor at Harvard Medical School, described a quantitative method that can correct such errors due to geographical ancestry known collectively as “population stratification”. It will help if the disease groups, sharing the same trait, have differences in their geographic ancestry.
Tags: Genome wide association studies, Genome-wide association studiesX, GWA, GWAS, Hap-Map project, HGP, human variome project, SNPs, variome
 No Comments »
No Comments »
Metagenomics is a culture independent approach that has contributed extensively to the study and understanding previously unidentified microbial communities. Seeking a further understanding of employing metagenomics in the study of the Red Sea microbial communities, we are pleased to interview Dr. Rania Siam, an Associate Professor in the Biology Department, the Director of the Biotechnology Graduate Program at the American University in Cairo (AUC) and an Investigator in the Red Sea Marine Metagenomics Project that is currently running at the AUC in collaboration with King Abdullah University of Science and Technology (KAUST), Woods Hole Oceanographic Institution and Virginia Bioinformatics Institute at Virginia Tech. Dr. Siam holds a Ph.D. in Microbiology and Immunology from McGill University. In addition, she held several post-doctoral positions at McGill Oncology Group, Royal Victoria Hospital, The Salk Institute for Biological Studies and The Scripps Research Institute. Since 2008, Dr. Hamza El Dorry (PI) and Dr. Siam (Co-PI) have been leading the Red Sea Metagenomics research team. The team is exploring novel bacterial communities in the Red Sea through actively participating in Red Sea expeditions for sampling and performing extensive molecular biology, genomics and computational analysis of the data.

1- Dr. Siam, thank you very much for accepting our request. Would you please explain for us the driving motives behind doing metagenomics research in the Red Sea?
The Red Sea is a unique environment in the region that remains to be explored. Thus, working on the Red Sea gives us the opportunity to perform essential research for the region. Furthermore, our main interest are the Red Sea brine pools that are unique environments in terms of high temperature, high salinity, high metal contents and low oxygen. Microbial communities living in these environments are known as extremophiles. The survival of extremophiles in such drastic conditions indicates their possession of genes with novel properties that underlie these unique survival characteristics. Thus, we are highly motivated to explore these novel microbial communities and their unique properties. In addition, we are interested in extracting biotechnological products from the Red Sea that can be beneficial as antimicrobials and anticancer agents.
2- Would you please outline for us the objectives of the project and the main activities inside and outside the laboratory?
In our project, one of our main aims is establishing a Red Sea marine genomic database to be accessed by scientists’ worldwide. Additionally, we are screening this database for biotechnological pharmaceutical products as enzymes and anticancer agents. There are three main activities in the project: sampling, molecular biology/genomics work and computational analysis of the data. Concerning sampling, it is a challenging process that requires rigorous planning where samples should be subjected to proper processing and storage till arrival to the labs. In labs, samples are subjected to different procedures starting from DNA extraction followed by Whole Genome Sequencing (WGS) to identify unknown genes or unique ones and help us understand novel microbial communities. This requires rigorous computational analysis to make sense of our data. Furthermore, we construct fosmid libraries for isolation and purification of genes of biotechnological interest as lipases and cellulases. In addition, we carry out 16s rRNA phylogenetic analysis on the microbial communities present in the samples.
3- Since the idea is novel, we would like to know about the nature of samples, the parameters and the challenges imposed during the process of sampling.
Basically, the sampling process requires well-equipped research vessels as the Woods Hole Oceanographic Institute ‘Oceanus’ and The Hellenic Center for Marine Research (HCMR) ‘Aegeo’. In addition, it is essential to have a team of physical oceanographics for adequate sampling.

http://www1.aucegypt.edu/publications/auctoday/AUCTodayFall09/Cure.htm
Image Source: AUC Today
We started with two different brine pools: Atlantis II Deep and Discovery Deep. As brine pools, these two regions are characterized by the presence of extremely harsh conditions as I mentioned before. In the 2008 and the 2010 KAUST Expedition to the Red Sea, we collected two forms of samples: Large volume water and sediments. In both cases, we face challenges during sample collection. Collecting large volume water samples can take up to 4 hours. In case of bad weather, it is nearly impossible to collect samples. Regarding the sediments, the heavy weight of the sediment core is the main challenge since it may drag people to the water during the sampling process.
The water samples are collected using CTDs (an acronym for Conductivity, Temperature and Depth), it is formed of 10 liter bottles that collect samples and measure the conductivity, temperature and depth, in addition to other parameters. CTDs are capable of measuring these physical properties from each meter of water. Accordingly, they are able to retrieve 2200 readings for each parameter at 2200 m depth. This is very beneficial as it allows us to correlate the physical and chemical parameters with the nature of microbial communities obtained from each sample.
4- What are the reasons behind the choice of Atlantis II Deep and Discovery Deep brine pools for study?
These sites are unique. Atlantis II Deep is 2200 meters below the water surface. It is characterized by having high temperature (68 °C), high heavy metal content, high salinity and little oxygen. Thus, these extremely drastic conditions are motivating us to explore the extremophilic microbial communities in this pool. Discovery Deep is adjacent to Atlantis II but the conditions are less harsh and varies in its heavy metal content. This encourages us to undergo comparative genomic analysis between the two regions.
5- Did the relatively new metagenomic approach of sequencing multiple genomes, combined with, the novelty of employing this approach in the study of microbial milieu in the Red Sea imply some unprecedented practical challenges in retrieving entire and authentic DNA sequences?
Yes, many challenges are present. For example, in case of sediments, many cells die. This in turn can make the process of DNA extraction more difficult. However, we managed to cope with this problem by quickly extracting the DNA from the sediments following its arrival to the labs and avoid freezing and thawing. Another problem with sediments is the presence of impurities that are co-extracted with DNA and interfere with the analysis. Concerning water samples, the main challenge is that the amount of DNA in the samples is very minute. This was dealt with by filtering large volumes of water up to 500 liters per sample.
6- Is recognizing and validating sets of data that are pointing out to specific patterns of microbial diversity or novel genes considered to be challenging?
Actually, we have retrieved enormous amount of data and a large percentage of the data has no match to sequences present in the other genomic databases. This has inspired us to think of new approaches for data analysis to identify the role of these novel sequences and seek collaborations with computational biologists.
7- Finally, do you think there are other environments in Egypt with unique properties which make it promising for employing metagenomics to discover novel genes and bacterial strains?
Yes, actually Egypt is very rich in environments with unique properties. For example, we have the deserts, Siwa‘s hot springs and the Nile. Many unique environments are yet to be explored and lots of research needs to be performed. Our natural resources are limitless.
Tags: 16s rRNA phylogenetic analysis, Aegeo, Atlantis II, AUC, Brine pools, Computational analysis, Computational biologists, CTDs, Culture-independent, Discovery Deep, DNA extraction, Drastic conditions, Expeditions, Extremophiles, Fosmid libraries, genomics, KAUST, Metagenomics, microbial communities, Molecular biology, Oceanus, Red Sea, Virginia Bioinformatics Institute, Virginia Tech., WGS, Woods Hole Oceanographic Institution
 2 Comments »
2 Comments »
 A 70-year-old man can enjoy the blessing of fatherhood to perfectly healthy children. That is practically a miracle quality-wise because the male gametes are being produced in huge amounts, 1000 per second to be exact. That amounts to about 30 billion a year. Each and every one of those sperms could potentially contribute by half to the formation of a human being. So how could this rapid production preserve and maintain the critical quality required? How could errors during the sperm production be avoided? Researchers have recently gained an insight into the molecular mechanism as released in the “Proceedings of the National Academy of Sciences”. A 70-year-old man can enjoy the blessing of fatherhood to perfectly healthy children. That is practically a miracle quality-wise because the male gametes are being produced in huge amounts, 1000 per second to be exact. That amounts to about 30 billion a year. Each and every one of those sperms could potentially contribute by half to the formation of a human being. So how could this rapid production preserve and maintain the critical quality required? How could errors during the sperm production be avoided? Researchers have recently gained an insight into the molecular mechanism as released in the “Proceedings of the National Academy of Sciences”.
During the sperm production, there is an automatic quality control process. This control mechanism is strengthened by a specific genetic addition, present in both humans and great apes. The triggering factor is comprised of parts of an endogenous retrovirus, incorporated in our genome. 15 million years ago, this viral DNA was presumably incorporated in the genetic makeup of one of our ancestors. A lucky coincidence? Perhaps, but the researchers claim it catched on during the process of evolution. The site of insertion of this viral DNA is close to a gene, responsible for the production of a crucial control factor.
The control factor, termed p63, drives faulty cells straight to their apoptosis. It imposes a strict quality control of the genome because even in cases of a slight damage to the DNA, the cells ultimately die. As a result, the passing on of a flawed genome to the next generation is prevented. Some cells definitely fall as victims in the process. In addition, this mechanism could protect against certain types of cancer as testicular carcinoma. They refer that, p63 represents a barrier to the formation of tumors in normal healthy tissues. In cases of testicular cancer, the administration of drugs, that restore the function of p63, could prove potentially useful in the near future.
Source: Pharmazeutische Zeitung and original PNAS paper (thanks to Mariam and Steve Moss for sharing)
Tags: evolution, p63, PNAS, quality control, retrovirus, sperm, testicular carcinoma, tumor
 3 Comments »
3 Comments »
How about taking a “closer” look onto a microbe??? I’ll try to take you on a journey deep into the nature of the particles constituting it, deep to the extent of subatomic levels, in a trial to learn more about the beginning of life and matter…
Since being a nuclear physicist has always been a dream for me, but obviously didn’t come true, I have been very interested lately in the news about the re-operation of the Large Hadrons Collider (LHC) at CERN and the experiments being conducted there in their attempts to find out more about the composition of matter and the origins of our universe, and the effects these discoveries will have on all different fields of science.
My readings into this topic have brought me to know more about one of the fundamental building blocks of matter, the Quark. Being an elementary particle, quarks theoretically can’t be broken down into smaller units. Their existence was first proposed by physicists Murray Gell-Mann and George Zweig in 1964 as “the Quark model”. The model was introduced to give a better explanation and understanding of atomic Nucleii composition, but there was little evidence for their existence. This lasted till “deep inelastic scattering “experiments were conducted at SLAC National Accelerator Laboratory operated by Stanford University in 1968. since then, six types of quarks also known as “the six flavors” have been discovered, divided into three generations : 1st generation including ( up) and (down) quarks, 2nd including (charm) and (strange) quarks and the 3rd including (bottom) and ( top) quarks. The (top) quark, first observed at “Fermilab” in 1995, was the last to be discovered. There have been trials to prove the existence of a 4th generation of quarks, but till now, all have failed but in the future, and thanks to the current LHC experiments, who knows? After all, protons, neutrons and even atoms were once considered fundamental units of matter and that there was nothing more beyond them!!!

Higher generations of quarks are heavier and less stable, so they undergo certain type of particle decay into the more stable types, those are the up and down quarks, the most abundant in our Universe. The higher generation quarks can’t be produced except at extreme conditions of heat and pressure and with the help of high energy collision, a state believed to exist just after “The big bang” that created our universe.
Unfortunately quarks can’t exist solely in space. They form composites known as Hadrons, the most stable of which are protons and neutrons (so quarks are the building units of the building units of the nucleus of an atom!). This is because of two very important physical phenomenons known as “color charge” and “strong interaction”. These very strong bonds are mediated by energy carriers known as “gluons” (actually derived the word glue, as to stick!) and therefore quarks can’t be isolated singularly, making their observation not an easy task for physicists through the years. Simulating the cosmic conditions present just after the big bang, at which quarks were supposed to exist singularly in what is known as “the quark-gluon plasma” is one of the major aims of the CERN experiments through the LHC.
So you might ask yourself, if they can’t be observed by themselves and can’t exist singularly under normal conditions, how can they prove their existence in the first place? The answer simply is that proposing their presence justifies a lot of physical phenomenons and fits into certain physical models and gives the right answers to lots of experiments, so science has to admit that they are there!
Quarks have lots of interesting characteristics. For example, they have fraction charges ( like -1/3, and +2/3). Each quark has an antiquark having the same magnitude, mean half life but opposite charges (+1/3, -2/3).Hadrons, consisting of quarks, will always have integer charges (for example, a neutron has a charge of 0 consisting of 1 up quark (+2/3) and 2 down quarks( 2*-1/3) that is a sum of zero). They are the only elementary particles in the Standard Model of particle physics to experience all four fundamental interactions, also known as fundamental forces (electromagnetism, gravitation, strong interaction, and weak interaction) .
The quarks got their name when Gell-Mann named them after the sound made by ducks!!!! The “strange” quarks were termed so because they had exceptionally long half lives!!.As for the “charm” type, Glashow, who co proposed charm quark with Bjorken, is quoted as saying, “We called our construct the ‘charmed quark’, for we were fascinated and pleased by the symmetry it brought to the subnuclear world.”
References:
http://en.wikipedia.org/wiki/Elementary_particle
http://www2.slac.stanford.edu/vvc/theory/quarks.html
http://public.web.cern.ch/public/en/lhc/ALICE-en.html
http://en.wikipedia.org/wiki/Quark
Tags: CERN, LHC, quarks, subatomic
 4 Comments »
4 Comments »
 On a Tuesday, the year 1995, Graham, a 57-year-old patient on the verge of congestive heart failure, received a call that a donor has been found. This donor committed suicide via a self-inflicted gunshot wound. After the successful transplant, Graham got in touch with the donor’s family, married his wife and after a whopping 12 years later, kills himself through the same technique. In an interview held a couple of years after his marriage, he admitted to the reporter that after seeing the donor’s ex-wife, he felt as if he had already known her for years. On a Tuesday, the year 1995, Graham, a 57-year-old patient on the verge of congestive heart failure, received a call that a donor has been found. This donor committed suicide via a self-inflicted gunshot wound. After the successful transplant, Graham got in touch with the donor’s family, married his wife and after a whopping 12 years later, kills himself through the same technique. In an interview held a couple of years after his marriage, he admitted to the reporter that after seeing the donor’s ex-wife, he felt as if he had already known her for years.
Ever since heart transplant surgeries were a success in 1967, scientists were skeptical about what is now referred to as, Cellular Memory Phenomenon. This was provoked through the close observation of recipients, who repeatedly report bizarre distant memories and new personal preferences. Exactly how much can the cells of an organ, other than the brain, mainly the heart in this case, store memories? The Discovery Channel aired a documentary titled “Transplanting memories” where various experts gave their opinion on the matter.
If such phenomenon truly exists, where are these memories located inside the cells? Could it be the DNA? But this is sheltered inside the nucleus and remains entangled except when cellular division takes place. This makes its access difficult, but after all, it cannot be THAT difficult, otherwise mutagenic agents wouldn’t have succeeded.
Possibly proteins. Dr. Candace Pert stated that, since the brain and human organs are linked through a massive network of peptides. She said “I believe that memory can be accessed anywhere in the peptide/receptor network. For instance, a memory associated with food may be linked to the pancreas or liver, and such associations can be transplanted from one person to another.”
Source: The Medical News
Image Source: Hiveworld
Tags: cellular memory, discovery channel, Graham, heart, phenomena, suicide, transplant, transplanting memories
 3 Comments »
3 Comments »
Not too long ago, I read about a research done at the Kennedy Institute of Rheumatology Division, which has identified a new ligand for Toll-like receptor 4. This receptor was previously known for activating the immune system through the detection of threats as lipopolysaccharide or gram-negative bacteria. The new ligand, Tenascin-C, is an extracellular glycoprotein, whose elevated expression in cases of inflammation provoked scientists to study its role in the process. 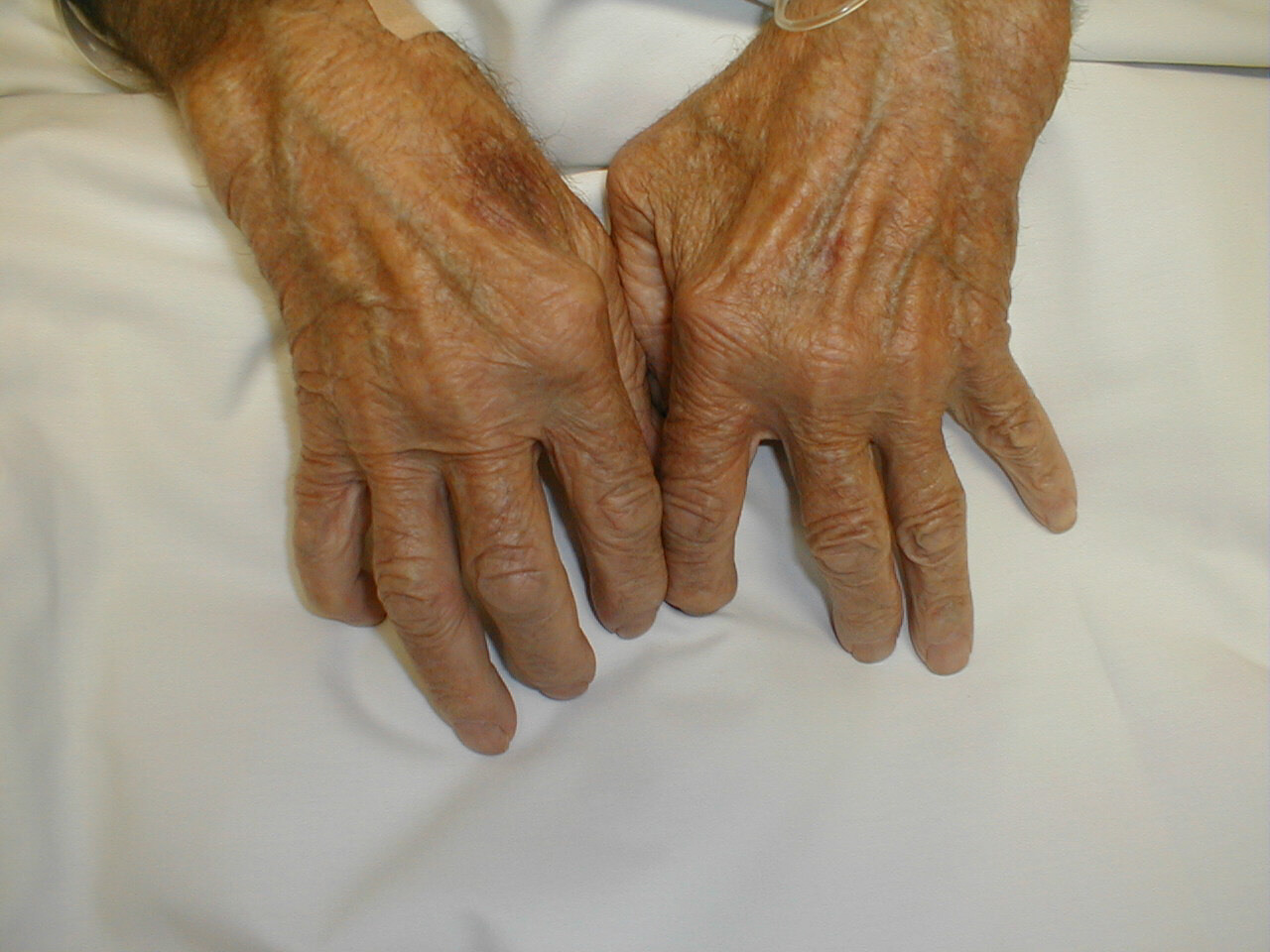
The study noted that its presence was critical to maintain the ongoing inflammation seen in cases of rheumatoid arthritis. In reference to this study, the author stated “We have uncovered one way that the immune system may be triggered to attack the joints in patients with rheumatoid arthritis. We hope our new findings can be used to develop new therapies that interfere with tenascin-C activation of the immune system and that these will reduce the painful inflammation that is a hallmark of this condition”
I was able to contact Dr Kim Midwood and obtained this brief interview:
1. Do you have any speculations as to why Tenascin-C is overly expressed in certain individuals causing prolonged inflammation cases, whilst remaining within normal levels in others?
What regulates tissue levels of tenascin-C is not currently known and this is something that we are working on finding out.
2. From the different ligands of TLR4, why was Tenascin-C of particular interest in your research?
I have a long standing interest in how cell behavior is influenced by the extracellular environment, and in particular the role of extracellular matrix proteins in regulating cell phenotype during the response to tissue injury. For the last 10 years, I’ve been studying the role of tenascin-C – a protein specifically and transiently expressed upon tissue injury, but persistently expressed in chronic inflammatory diseases such as rheumatoid arthritis. This pattern of expression, plus the high homology of tenascin-C domains to other known pro-inflammatory matrix molecules or ‘DAMPs’ prompted us to investigate whether tenascin-C was an endogenous activator of the immune response and whether its persistent expression in RA contributed to disease pathogenesis.
3. What do you think the extent of similarity will be between the mice & human response to the Tenascin-C blockage?
I cannot predict how differently the mouse and human will behave.
4. Do you suspect a certain mechanism of the increase in inflammatory molecules caused by Tenascin-C?
We know that tenascin-C activates TLR4, activation of this receptor is well known to induced the expression of pro-inflammatory genes via activation of many intracellular signaling pathways.
5. How do you see the potential of such study for rheumatoid arthritis patients?
We plan to identify ways to inhibit the pro-inflammatory action of tenascin-C in the hope that this may be useful in reducing chronic inflammation in the joint.
Original research paper: Tenascin-C is an endogenous activator of Toll-like receptor 4 that is essential for maintaining inflammation in arthritic joint disease. Nature Medicine 15, 774 – 780 (2009). PMID: 19561617 (Vote for the abstract on Biowizard)
Image Credit: Davidson College Undergraduate Course
Tags: chronic inflammation, DAMPs, endogenous, inflammation, Interview, intracellular signal, Kim Midwood, ligand, pro-inflammatory, rheumatoid arthritis, rheumatoid therapy, Tenascin, TLR4, Toll-like receptor
 No Comments »
No Comments »
When you hear/ read the term “Phage Therapy“, you’ll be automatically directed to the concept of using bacteriophages, the virus-like particles that infect bacteria, to kill/ lyse the resistant bacterial strains, instead of the “useless” antibiotics that allowed bacteria to fool them & develop resistance against them. The initial target of phage therapy was to kill the bacteria using phages; because they act like any other virus; get in, multiply and lyse the cell. But, by this way, bacteria develop resistance against phages more rapidly. So, they may become useless by time. In this paper from PNAS: “Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy,” two bioengineers, Timothy K. Lua and James J. Collins, from Boston University successfully engineered the Enterobacteria filamentous phage M13 to weaken bacteria not to kill it. Sounds strange, right? By engineering M13, they gave us a variety of options:
1st, we may make M13 overexpress a bacterial protein named lexA3 which inhibits the ability of the bacteria to repair their damaged DNA by the action of Ofloxacin –as pharmacophils, who had 2 consecutive chemotherapeutics courses, we may recall that quinolones’ MOA is generation of ROS. So, the repressor suppresses the bacterial SOS mechanism. Very promising results were observed; the adjuvant therapy increased the survival rate of mice infected with resistant E. coli. It was also observed that the adjuvant therapy reduced the rate of developing mutations/ resistance within the E. coli population.

2nd, bacteriophage can be responsible for expression of certain proteins that can attack gene networks in bacteria which are not target for existing antibiotic classes. I will mention just one example here, expression of CsrA which is a “global regulator of glycogen synthesis and catabolism, gluconeogenesis, and glycolysis, and it also represses biofilm formation,” biofilms is thought to be related to antibiotic-resistance and OmpF porin which is used by quinolones to enter the bacterial cell, it may enhance its entrance.

Now, thanks to the engineered phages, we can use the old beloved antibiotic classes to treat bacterial infection using the engineered phages as an adjuvent therapy to potentiate the cidal action of the antibiotic on the former-resistant strains. A precaution was made to ensure that no lysogeny would take place in the human cells is that the phages were engineered to be “nonreplicative”. But we still have two problems regarding Phage Therapy in general: identifying the strain responsible for the infection & making sure that the human immune system won’t elicit an immune response against phages, they’re “foreigners” after all!
Image credits:
1- “Schematic of combination therapy with engineered phage and antibiotics. Bactericidal antibiotics induce DNA damage via hydroxyl radicals, leading to induction of the SOS response. SOS induction results in DNA repair and can lead to survival. Engineered phage carrying the lexA3 gene (lexA3) under the control of the synthetic promoter PLtetO and an RBS acts as an antibiotic adjuvant by suppressing the SOS response and increasing cell death”: http://www.pnas.org/content/106/12/4629.figures-only
2- “CsrA suppresses the biofilm state in which bacterial cells tend to be more resistant to antibiotics. OmpF is a porin used by quinolones to enter bacterial cells. Engineered phage producing both CsrA and OmpF simultaneously (csrA-ompF) enhances antibiotic penetration via OmpF and represses biofilm formation and antibiotic tolerance via CsrA to produce an improved dual-targeting adjuvant for ofloxacin”: http://www.pnas.org/content/106/12/4629.figures-only
Tags: antibiotic, bacteriophage, bioengineering, engineered bacteriophage, M13, ofloxacin, phage, phage therapy, resistant
 2 Comments »
2 Comments »
 On February 28, 1953, James Watson and Francis Crick announced to their friends that they have discovered the chemical structure of the DNA. After publishing their paper in Nature on April 2, the official announcement took place on April 25. On February 28, 1953, James Watson and Francis Crick announced to their friends that they have discovered the chemical structure of the DNA. After publishing their paper in Nature on April 2, the official announcement took place on April 25.
That is what I have read in Al-Ahram newspaper today. What a discovery! Imagine if they had not done it, we would have had no clue about genes & protein synthesis, no recombinant DNA tech- & no sequencing. We would have had no molecular biology departments in universities! Abby from NCIS & Greg from CSI would have had no job!
So, what was the real story? To what extent are the “rumors” saying that Rosalind Franklin is the real discoverer of the DNA double helix right? Is she really the “Dark Lady of DNA”?
I hope we can get the story through your comments after we read this:
1- Crick papers from the National Library of Medicine.
2- Watson’s interview with a group of top North Carolina high school students in 2003.
3- BBC celebrating the 50th anniversary of DNA structure discovery in 2003. (really interesting)
4- Rosalind Franklin: Dark Lady of DNA by NPR (National Public Radio)
Image credits:
Watson – Crick DNA model: http://www.cs.princeton.edu/
Rosalind Franklin: http://www.npr.org/
Tags: 1408f278d2a6f884de58c0c44b7a6af7, DNA, Francis Crick, James Watson, Rosalind Franklin
 No Comments »
No Comments »
Hello, hello. You’re now tuned to your favorite blog: micro-writers.egybio.net. Tonight we have this very special guest, live, online. After two months of waiting, we finally got this exclusive interview with the emerged Streptococcus pyogenes strain, the most dangerous ever, M1T1. We have it here, with us, in the studio.
– Hello, M1T1. Welcome in our studio.
– Hey there.
– We knew from our resources, which are totally classified, that you got yourself in trouble recently.
– (Interrupting), I did NOT get myself in trouble. EID set me up.
– M1T1, Would you please calm down & tell us a little more about yourself?
– Well, I belong to Group A streptococci (GAS) aka Streptococcus pyogenes. M1T1 is my serotype; I’m just a clonal strain. As you know, S. pyogenes colonize human skin & throat causing either non-invasive (sore throat, tonsillitis & impetigo) or invasive (necrotizing fasciitis NF, scarlet fever & streptococcal toxic-shock syndrome STSS) infections. Actually, NF gave me my nick: Flesh-eating bacteria.
– So, you cause all people NF & STSS?
– No, kid. It depends on their genetic susceptibility, what you call “Host–pathogen interactions”. I was isolated from patients with invasive as well as non-invasive infections during 1992–2002. This is NOT entirely my fault; humans can make me extra virulent by selecting the most virulent members.
– Back to your history, when have you exactly been isolated?
– M1 & her sisters were the worst nightmare in US & UK in the 19th century as they caused the famous pandemic of scarlet fever. “Nevertheless”, early 1980s was the golden age of my strain as well as my very close sisters M3T3 & M18. We caused STSS & NF in different parts of the world. Great times, great times!
– Only for you, I suppose! So, what made you hypervirulent? What caused you this “epidemiologic shift”?
– Two reasons Dr Ramy K. Aziz identified that improved my fitness to humans: the new genes I got from phages & “host-imposed pressure”. Both resulted in the selection & survival of me M1T1 the hypervirulent strain. Dr Aziz’s work at Dr Kotb’s lab resulted in identification of a group of genes I got from phages that changed my entire life.
– Interesting! Tell us more about that. How did phages “change your life”?
– Dr Aziz proved that I differ from my ancestral M1 when he found that I have 2 extra prophages (lysogenized phages didn’t get the chance to lyse me, so they became integrated in my genome):
1. SPhinX which carries a gene encodes the potent superantigen SpeA or pyrogenic exotoxin A (scarlet fever toxin).
2. PhiRamid which carries another gene encodes the most potent streptococcal nuclease ever, Sda1.
3. He also found that phages conversion from the lytic state to the lysogenic state resulted in exchange of toxins between our different strains (aka Horizontal Gene Transfer). Phages are very good genetic material transporters, what makes “strains belonging to the same serotype may have different virulence components carried by the same or highly similar phages & those belonging to different serotypes may have identical phage-encoded toxins.” What a quote from Rise and Persistence of Global M1T1 Clone of Streptococcus pyogenes.
– Well, It was not that interesting. So, what? What’s the significance? How that made you hypervirulent?
– You can’t get it? You’re not that smart, are you? Tell me, what made M1 hypervirulent causing scarlet fever in the 1920s and me hypervirulent causing STSS in the 1980s with a 50-years decline period?
– Superantigen?
– Exactly. You do have your moments! Superantigen encoding-gene was present in us and absent in strains isolated in the period between them. The interesting part, for me of course, that humans after 50 years of absence of hypervirulent strains had absolutely no superantigen-neutralizing antibodies. That was the real invasive party. Superantigen causes high inflammatory response because of its non-specific binding to immune system components (antibodies & complements) causing an extremely high inflammatory response. In fact, SuperAg inflammatory response is “host-controlled”.
– So, what about Sda1?
– Streptodornase (streptococcal extracellular nuclease) helps me to degrade neutrophils that entrap me in the neutrophil extracellular traps (NETs). So, I can invade humans freely & efficiently and be able to live in their neutrophils. Dr Aziz proved in his paper “Post-proteomic identification of a novel phage-encoded streptodornase, Sda1, in invasive M1T1 Streptococcus pyogenes” that it’s all about C-terminus in my Sda1; the frame-shift mutation increased my virulence while deletion decreased it.
– Now we know about your SuperAg & nuclease (DNase), what’s the “host-imposed pressure”?
– I have my own SpeB (Protease), I use it to degrade my other proteins (virulence factors), which provides me with a good camouflage & gives me access to blood. When the host immune system recognizes me, it traps me in NETs. At this time, I secret Sda1 to degrade neutrophils. Actually, SpeB protects you, humans, from my Sda1& my other toxins. When SpeB was compared in patients with severe & non-severe strep infections, it was found that SpeB wasn’t expressed in case of severe infection. Expression of SpeB may be host-controlled, as host selects the mutants with a mutation in covS, a part of my regulatory system which regulates my gene expression including SpeB gene.
– Finally, M1T1. How do you see your future?
– More new phage-encoded genes, more selection of the hypervirulent strains by the host & more regulation of expression of my virulence factors. Pretty good future! I also count on humans to not develop immunity against me like what happened in 1980, when I got new virulence factors or allelic variations in my old ones.
Thank you, M1T1. Pleasure talking to you…….M1T1? M1T1, where are you? Why do I feel this strange pain in my throat?
Image credits:
Streptococcus pyogenes: http://adoptamicrobe.blogspot.com/
Tags: covS, epidemiologic shift, flesh-eating bacteria, global M1T1 clone, group A streptococci (GAS), horizontal gene transfer, host-imposed pressure, host–pathogen interactions, impetigo, M1T1, Malak Kotb, necrotizing fasciitis, neutrophil extracellular traps (NETs), phage-encoded toxins, pyrogenic exotoxin A, Ramy K. Aziz, S. pyogenes, scarlet fever, sore throat, streptococcal nuclease Sda1, streptodornase, STSS, superantigen SpeA, tonsillitis
 5 Comments »
5 Comments »
|
















 Entries (RSS)
Entries (RSS)