Author Archive
Metagenomics is a culture independent approach that has contributed extensively to the study and understanding previously unidentified microbial communities. Seeking a further understanding of employing metagenomics in the study of the Red Sea microbial communities, we are pleased to interview Dr. Rania Siam, an Associate Professor in the Biology Department, the Director of the Biotechnology Graduate Program at the American University in Cairo (AUC) and an Investigator in the Red Sea Marine Metagenomics Project that is currently running at the AUC in collaboration with King Abdullah University of Science and Technology (KAUST), Woods Hole Oceanographic Institution and Virginia Bioinformatics Institute at Virginia Tech. Dr. Siam holds a Ph.D. in Microbiology and Immunology from McGill University. In addition, she held several post-doctoral positions at McGill Oncology Group, Royal Victoria Hospital, The Salk Institute for Biological Studies and The Scripps Research Institute. Since 2008, Dr. Hamza El Dorry (PI) and Dr. Siam (Co-PI) have been leading the Red Sea Metagenomics research team. The team is exploring novel bacterial communities in the Red Sea through actively participating in Red Sea expeditions for sampling and performing extensive molecular biology, genomics and computational analysis of the data.

1- Dr. Siam, thank you very much for accepting our request. Would you please explain for us the driving motives behind doing metagenomics research in the Red Sea?
The Red Sea is a unique environment in the region that remains to be explored. Thus, working on the Red Sea gives us the opportunity to perform essential research for the region. Furthermore, our main interest are the Red Sea brine pools that are unique environments in terms of high temperature, high salinity, high metal contents and low oxygen. Microbial communities living in these environments are known as extremophiles. The survival of extremophiles in such drastic conditions indicates their possession of genes with novel properties that underlie these unique survival characteristics. Thus, we are highly motivated to explore these novel microbial communities and their unique properties. In addition, we are interested in extracting biotechnological products from the Red Sea that can be beneficial as antimicrobials and anticancer agents.
2- Would you please outline for us the objectives of the project and the main activities inside and outside the laboratory?
In our project, one of our main aims is establishing a Red Sea marine genomic database to be accessed by scientists’ worldwide. Additionally, we are screening this database for biotechnological pharmaceutical products as enzymes and anticancer agents. There are three main activities in the project: sampling, molecular biology/genomics work and computational analysis of the data. Concerning sampling, it is a challenging process that requires rigorous planning where samples should be subjected to proper processing and storage till arrival to the labs. In labs, samples are subjected to different procedures starting from DNA extraction followed by Whole Genome Sequencing (WGS) to identify unknown genes or unique ones and help us understand novel microbial communities. This requires rigorous computational analysis to make sense of our data. Furthermore, we construct fosmid libraries for isolation and purification of genes of biotechnological interest as lipases and cellulases. In addition, we carry out 16s rRNA phylogenetic analysis on the microbial communities present in the samples.
3- Since the idea is novel, we would like to know about the nature of samples, the parameters and the challenges imposed during the process of sampling.
Basically, the sampling process requires well-equipped research vessels as the Woods Hole Oceanographic Institute ‘Oceanus’ and The Hellenic Center for Marine Research (HCMR) ‘Aegeo’. In addition, it is essential to have a team of physical oceanographics for adequate sampling.

http://www1.aucegypt.edu/publications/auctoday/AUCTodayFall09/Cure.htm
Image Source: AUC Today
We started with two different brine pools: Atlantis II Deep and Discovery Deep. As brine pools, these two regions are characterized by the presence of extremely harsh conditions as I mentioned before. In the 2008 and the 2010 KAUST Expedition to the Red Sea, we collected two forms of samples: Large volume water and sediments. In both cases, we face challenges during sample collection. Collecting large volume water samples can take up to 4 hours. In case of bad weather, it is nearly impossible to collect samples. Regarding the sediments, the heavy weight of the sediment core is the main challenge since it may drag people to the water during the sampling process.
The water samples are collected using CTDs (an acronym for Conductivity, Temperature and Depth), it is formed of 10 liter bottles that collect samples and measure the conductivity, temperature and depth, in addition to other parameters. CTDs are capable of measuring these physical properties from each meter of water. Accordingly, they are able to retrieve 2200 readings for each parameter at 2200 m depth. This is very beneficial as it allows us to correlate the physical and chemical parameters with the nature of microbial communities obtained from each sample.
4- What are the reasons behind the choice of Atlantis II Deep and Discovery Deep brine pools for study?
These sites are unique. Atlantis II Deep is 2200 meters below the water surface. It is characterized by having high temperature (68 °C), high heavy metal content, high salinity and little oxygen. Thus, these extremely drastic conditions are motivating us to explore the extremophilic microbial communities in this pool. Discovery Deep is adjacent to Atlantis II but the conditions are less harsh and varies in its heavy metal content. This encourages us to undergo comparative genomic analysis between the two regions.
5- Did the relatively new metagenomic approach of sequencing multiple genomes, combined with, the novelty of employing this approach in the study of microbial milieu in the Red Sea imply some unprecedented practical challenges in retrieving entire and authentic DNA sequences?
Yes, many challenges are present. For example, in case of sediments, many cells die. This in turn can make the process of DNA extraction more difficult. However, we managed to cope with this problem by quickly extracting the DNA from the sediments following its arrival to the labs and avoid freezing and thawing. Another problem with sediments is the presence of impurities that are co-extracted with DNA and interfere with the analysis. Concerning water samples, the main challenge is that the amount of DNA in the samples is very minute. This was dealt with by filtering large volumes of water up to 500 liters per sample.
6- Is recognizing and validating sets of data that are pointing out to specific patterns of microbial diversity or novel genes considered to be challenging?
Actually, we have retrieved enormous amount of data and a large percentage of the data has no match to sequences present in the other genomic databases. This has inspired us to think of new approaches for data analysis to identify the role of these novel sequences and seek collaborations with computational biologists.
7- Finally, do you think there are other environments in Egypt with unique properties which make it promising for employing metagenomics to discover novel genes and bacterial strains?
Yes, actually Egypt is very rich in environments with unique properties. For example, we have the deserts, Siwa‘s hot springs and the Nile. Many unique environments are yet to be explored and lots of research needs to be performed. Our natural resources are limitless.
Tags: 16s rRNA phylogenetic analysis, Aegeo, Atlantis II, AUC, Brine pools, Computational analysis, Computational biologists, CTDs, Culture-independent, Discovery Deep, DNA extraction, Drastic conditions, Expeditions, Extremophiles, Fosmid libraries, genomics, KAUST, Metagenomics, microbial communities, Molecular biology, Oceanus, Red Sea, Virginia Bioinformatics Institute, Virginia Tech., WGS, Woods Hole Oceanographic Institution
 2 Comments »
2 Comments »
Hansenula polymorpha, also known as Pichia angusta, with its metabolism highly dependent on methanol as a carbon source, has been excessively employed in the production of therapeutic proteins for the last two decades. The yeast was first discovered in 1950s in spoiled orange juice.

Image source:
http://www.drugdevelopment-technology.com/contractors/contract_research/artes/
The biotechnological interest in Hansenula is mainly attributed to its unique capabilities underlied by its rare characteristics. For instance, being one of the limited group of yeast that are able to assimilate methanol, gives it the advantage of being able to utilize relatively cheap substrates. In addition, Upon high temperature, Hansenula polymorpha shifts its biochemical methanol metabolism pathway to the biosynthesis of trehalose which is a thermo-protective sugar, this fact explains its unique ability to resist temperatures up to 49 degree Celsius. further, Hansenula is able to secrete the protein products directly to the culture, a fact that renders the whole process of downstream processing easier and less costly. finally, the ability to survive in wide pH range,from 2.5 to 6.5, makes it a versatile protein factory which is exploited in the production of various types proteins, each of which requires a very different optimum pH value throughout the fermentation process.

Image source:
http://en.wikipedia.org/wiki/Hansenula_polymorpha
( Budding Hansenula cells)
However, with Hansenula polymorpha post-translational modification processes are not highly regulated, this makes Hansenula useful for the production of relatively small to medium sized polypeptides as Parathyroid hormone, Staphylokinase, Elafin……etc. However, Regarding large polypeptides, mammalian cells with tightly regulated post-translational modification processes represent a better option.
On the industrial scale, different strains of Hansenula polymorpha are being exploited as expression systems. For instance, a genetically engineered strain known as super-transformed strain bearing additional two advantages than the wild type strain. Being super-transformed means that it is capable of secreting Calnexin which is a protein Chaperone that functions to ensure proper folding of the secreted protein. Additionally, Calnexin enhances the protein secretion efficiency of Hansenula . Accordingly, the super-transformed strain gains its industrial potential in terms of quality assurance of the secreted protein product as well as increasing the cost effectiveness of the industrial process.
References: 1.“Hansenula polymorpha” wikipedia, www.wikipedia.org
2. Gellissen G, “Hansenula ploymorpha : Biology & Applications”, 2002
3.Marcos A. Oliveira, Victor Genu, Anita P.T. Salmazo, Dirce M. Carraro and Gonçalo A.G. Pereira1 ”The transcription factor Snf1p is involved in a Tup1p-independent manner in the glucose regulation of the major methanol metabolism genes of HansenulaPolymorpha”, Genetics and Molecular Biology, 521-528 (2003)
4. Giuseppinia Parpinello, Enrico Berardi, and Rosanna Strabbioli,” A Regulatory Mutant of Hansenula polymorpha Exhibiting Methanol Utilization Metabolism and Peroxisome Proliferation in Glucose”, J Bacteriol. 1998 June; 2958–2967.
Tags: Calnexin, Chaperone, Downstream processing, Elafin, Hansenula polymorpha, methanol, Parathyroid hormone, Post-translational modification, Staphylokinase, Super-transformed, Therapeutic proteins, thermo-protective, trehalose, Yeast
 No Comments »
No Comments »
Despite great advances in cancer therapy, conventional therapies are still implementing drawbacks and dilemmas that drives cancer research to consider other strategies to overcome the drawbacks implicated by conventional cancer therapy. The cancer treatment using anticancer agents possesses many adverse events related to bone marrow suppression and death of other rapidly proliferating cells is resulting from the either of the two following reasons:
- The narrow therapeutic index of anticancer agents that is in many cases hard to adjust with the inter-individual Pharmacokinetic &/or pharmacogenetic variation among cancer patients.
- The lack of specificity in most of anticancer agents systemically administered where anticancer agents kill both tumor cells and healthy cells.
Both of the above facts driven cancer therapy research in to many different aspects in the hope of optimizing the therapy & providing new techniques to get over the drawbacks of conventional therapy; The pharmacokinetics & /or pharmacogenetics, gene therapy in addition to the targeting therapy including the nanotherapy.

Professor Mostafa El Sayed was awarded the 2007 US National Medal of Science for his huge contribution in the field of nanotherapy in cancer as a molecular targeting approach that overcomes side effects of conventional cancer therapy. The idea lies in two main aspects: The first is molecular targeting & the second is photothermal destruction of malignant cells. The technique encompasses injecting gold nanoparticles conjugated with anti- Epidermal Growth Factor Receptor “anti -EGFR” monoclonal antibody , where the anti-EGFR is responsible for the the specific targeting which is molecularly based on the fact that epithelial carcinoma cells, particularly over-expresses “EGFR”. Regarding the tumor cidal effect it is mainly dependent on the photothermal destruction which is the role of the laser beam & gold nanoparticles, where particular nano size of gold makes it able to absorb light in Near Infra Red Region , which is the region where optical penetration is optimal & scatter laser beam and convert the light energy to thermal energy that is able to damage cell membrane and release the digestive enzymes and hence the death of cancer cells.
Image credit: www.gatech.edu
The choice of the laser light is a matter of the cancer location , where in case of cancer under the skin, Near Infra Red “NIR” laser light is recommended for its larger penetration depth. Gold particles are especially used because they are easily bioconjugated & they served as photoabsorbers due to overlap of absorption band of their specific nanosize with with argon laser beam. The gold nanoparticles are having a silica core and a gold shell & their absorption in the NIR is tuned by adjusting gold layer thickness as well as the size of silica core.
The pioneering success of the technique has been proved effective upon accumulation of Anti-EGFR antibody conjugated gold nanoparticles selectively in carcinoma cells and survival of benign cells as demonstrated by microscopic pictures of both benign & cancer cells as follows:
Gold nanoparticles are concentrated in cancer cells. “Pic 1”
Image credit: www.gatech.edu

Gold nanoparticles are not retained in benign cells. “Pic 2”
Image credit: www.gatech.edu
References:
- Ivan H. El Sayed, Xiaohua Huang, Mostafa A. El Sayed, ” Selective laser Photo-thermal therapy of epithelial carcinoma using anti-EGFR antibody conjugated gold nanoparticles”, Cancer Letters 239 (2006) 129-135.
- Erin B. Dickerson, Erick C. Dreaden, Xiaohua Huang, Ivan H. El Sayed,Hunghao Chu, Sujatha Pushpanketh, John F. Mcdonald, Mostafa A. El Sayed, ” Gold nano assiated near-infrared Plasmonic Phototheramal Therapy (PPTT) of squamos cell carcinoma in mice”, Cancer Letters 269 ( 2008) 57- 66.
Tags: adverse events, anticancer, argon laser, bone marrow suppression, cancer, Epithelial Carcinoma, Gene Therapy, laser, Molecular targetting, monoclonal antibody, nanotherapy, narrow therapeutic index, Pharmackinetic, pharmacogenetic, Photo-Thermal Destruction, Professor Mostafa El Sayed
 4 Comments »
4 Comments »
Normally when we hear the word immunity, we think of defense against infections, graft rejection, inflammation, etc. But, what if this defense may cause more damage than the infection itself ? In this case, the immune system shows a certain privilege through acting smarter where it deviates its mechanisms in a way to down-regulate its own damaging mechanisms. This whole process is known as Anterior Chamber-Associated Immune Deviation (ACAID).

image credit: http://research.opt.indiana.edu/
ACAID is endogenous to specific sites such as the anterior chamber of the eye and the brain. The process is considered as saving in cases of ocular infections where the visual axis is easily deflected by inflammation leading to blindness. Also ACAID is considered beneficial in case of allografting as it downregulates the immune processes responsible for allograft rejection. The process was discovered by Medwar in 1940, when he first noticed that surprisingly certain tumors proliferate more rapidly in the anterior chamber of the eye than anywhere else. Medwar’s further studies demonstrated the role of the process in transplantation immunology.
As an example of ACAID, upon antigenic inoculation of anterior chamber of the eye, the immune deviation presents itself as follows: Instead of Natural Killer T-cells perform certain functions attributed to T-Helper & T-Cytotoxic cells, the antigen injected into the eye APCs (Antigen Presenting Cells) that carry antigen to the spleen. These APCs activate NKT cells which in turn produce certain cytokines as TGF-ß that induce the generation of CD8+Tr cells which by production of cytokines such as TGF-ß and IL-10, can downregulate subsequent Th1-mediated DTH reactions against the same antigen.

image credit : http://www.nature.com
Thus, regarding the beneficial effects that can be drawn from ACAID, current research is being conducted for inducing ACAID to avoid graft and transplant rejection. ACAID can be induced by animal injection with non-ocular APCs, e.g., peritoneal exudate cells (PECs) that have been precultured with TGF-ß and antigen in vitro. Such procedure is believed to be a step forward toward the success of transplantation.
Tags: Anterior Chamber Associated Immune Deviation, APC, cytokines, graft, immunity, Medwar, ocular, T cells, Visual axis
 No Comments »
No Comments »
As we learned in pharmacology, the drug upon reaching the site of action, it gives a certain response. However, what if that response varies among individuals in terms of potency, duration or even adverse effects. As an example, in 1950s it has been noticed that Caucasians show prolonged effects to suxamethoniun chloride where further investigations revealed that 1 in every 3500 Caucasians has a less efficient butyryltranseferase, that metabolizes suxamethonium chloride, thus showing prolonged half-life & slower recovery from surgical paralysis. This fact has led to the emergence of a very unfavourable term “especially to clinicians”, Idiosyncrasy, which means the abnormal response to drugs, food & toxic agents that is peculiar to an individual. However, understanding the basic underlying mechanisms that led to this variation, made the scientists realize that in this case, the drug response is not only a matter of pharmacodynamic effects but is considered as a ring holding pharmacology at one side and the genetic make up (which is the DNA sequence of any protein dealing with the drug as receptors, carriers, metabolizing enzymes………, etc) at the other side. This understanding led the scientists to replace the old term of Idiosyncrasy with a better descriptive term known as pharmacogenetics.

image credit: www.medigenomix.de/
Variations in the genetic make up are diverse in types. The most common type is Single Nucleotide Polymorphisms (SNPs) which are single nucleotide substitutions that can be found in coding and non-coding regions in the DNA sequences of protein systems dealing with the drug. Another form of variation can be chromosomal aneuploidy as Trisomy, where an extra copy of a chromosome, carrying a gene coding for any protein system dealing with the drug, will in turn lead to response variation. As an example for that is in leukemic patients with down syndrome, since there is a third copy of chromosome 21 which bears Reduced folate Carrier gene that codes for the transmembrane Reduced Folate carrier system which is responsible for the of transport methotrexate inside the cell, thus having a third extra copy of this gene leads to the over expression of RFC & methotrexate toxicity due to high levels of intracellular methotrexate.
Now, with the fact that the genetic make up of an individual dictates the response has been laid down, the clinical application has become a consequent step. For example, regarding leukemic patients having down syndrome, as mentioned previously, they are highly predisposed to methotrexate toxicity & thus as a routine, these patients should be set on lower doses of methotrexate. This particular example is a little bit simple that is easily characterized by the well known phenotype of down syndrome patients. However, translating the basic knowledge of pharmacogenetics to useful clinical guidelines is a more complicated approach in terms of both characterizing the alteration in the genetic make up and the subsequent clinical decision regarding the choice of the drug, dose and the dosing schedule. For instance carrying out a SNP Genotyping for DNA sequencing and identification of SNPs is a difficult matter as SNPs are very scattered along the human genome, where it has been estimated that every 1,ooo base pair, one SNP is found. In addition, the functional characterization of this SNP on the protein level is a matter of complexity that requires well controlled studies (i.e. all other causes of response variation regarding the drug of interest are eliminated).

Inspite of the complexity of the investigations for clinical application, it is a productive promising approach that does have three positive impacts on the clinical application where upon the tailoring of the treatment protocol according to each patient genetic make up (individualized therapy), this will increase the efficiency of the medications, decrease side effects, adverse drug reactions, morbidities Image credit:& mortalities and thus reduce the finances of the clincal mangement of adverse drug reactions as well as toxicities that may be result from incompatibility of the drug with the genetic make up of the patient.
http://www.pharmacogeneticsinpsychiatry.com
Read the rest of this entry »
Tags: aneuploidy, Clinical application, functional characterization, Genetics, idiosyncrasy, pharmacogenetics, Reduced Folate Carrier, response variation, SNPs, suxamethonium chloride, trisomy
 No Comments »
No Comments »
Many genes have the same function whether they are inherited from the father or the mother. However, few genes are active only when they are inherited from mother & others are active only when they are inherited from father. This fact makes us raise our hands with a few questions: when we recieve genes of same function from both parents, which one’s action will predominate? and what are the basis of this selective predomination of action? Here comes the rule of what is known as “Genetic imprinting” where certain genes inherited from a certain parent will be silenced by an epigenetic mechanism rendering them inactive. This process happens mainly during the development of gametes.
The fraction of imprinted genes in the human genome is still unknown, however studies refer that 10%-25% of mouse genome is imprinted.
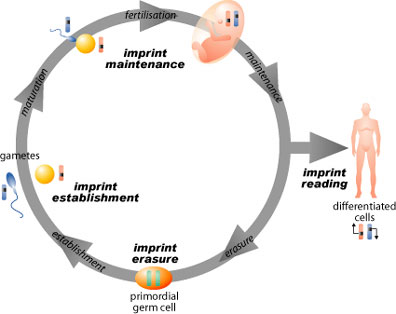
image credit:
www.bioteach.ubc.ca
Imprinted genes are localized in certain clusters in the genome where the whole cluster is silenced by methylation through certain methylases that act mainly by addition of methyl group to cytosine of spesific CpG dinucleotides within the clusters, so reducing the expression of the rest of genes in the clusters. However, Genetic imprinting is liable to modification along generations, for example if a male recieved imprinted genes from his mother, if it happens that this male will have a daughter, these imprinted genes he recieved from his mother will be activated by another process known as acetylation, where acetylated genes are actively tarnscribed. Thus, whether genes are imprinted or not depends only whether they came from a mother or father & is not a trait being passed through generations ( i.e. Non Mendelian pattern of inheritance). Diseases and developmental disorders are associated mainly when ceratin genes fail to be imprinted. Cancers have deen correlated with failure to imprint growth factors.
Tags: acetylation, Genetic, Imprinting, methylation, non mendelian
 2 Comments »
2 Comments »
Mitochondria contain their own DNA which is about 17,000 bp. Mutation in mtDNA can lead to a series of disorders known as mitochondrial disorders which will be very vivid in organs requiring high energy supply as brain, heart, eye & skeletal muscles. Owing to its oxidative function, mitochondria are having a high rate of mutations estimated to be 10 times that of nuclear DNA . However, mitochondrial disorders result also from mutations in nuclear DNA in the regions coding for mitochondrial components . For individuals having mitochondrial disorders resulting from mtDNA mutations , they have a mixture of mutated mtDNA & normal mtDNA which is a pattern of distribution known as mitochondrial DNA heteroplasmy. The proportion as well as the distribution of defective mtDNA influence the location , onset as well as severity of diseases.

image credit: www.mrc-dunn.cam.ac.uk
Conventional treatment of mitochondrial disorders is mainly supportive where patients are treated with co-enzyme Q10, which is a cofactor required for electron transfer from complexes I & II to complex III. However Gene Therapy & Modern Targetting Systems stepped in offering a radical & permenant cure for mitochondrial disorders. One of most promising strategies of Gene Therapy is allotropic expression which involves the engineering of normal genes then their introduction to nucleus. This method showed success in a mitochondrial disorder characterized by respiratory deficient phenotypes caused by a mutant MATP8 gene , where the engineered ATP gene has succeeded in production of ATPase 8 protein.
Modern targetting systems have developed offering a unique solution to mitochondrial disorders & overcoming the dilemma of heteroplasmy as well. Cell membarne Crossing Oligopolymers (CMCO’s) are molecules that are capable of penetrating cell membrane & selectively bind to mutated mtDNA inhibiting its replication. This technique has been successful in handling Oxidative Phosphorylation deficiency (OXPHOS deficiency).
Tags: allotropic expression, CMCO's, heteroplasmy, mitochondria, mtDNA
 No Comments »
No Comments »
|
















 Entries (RSS)
Entries (RSS)