Archive for the “Bacteriology” Category
Dr. Betsey Dexter Dyer is a professor of Biology at Wheaton College, Massachusetts, USA. She received her Ph.D.  from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including
The Modern Scholar: Unseen Diversity: The World of Bacteria (Audiobook), The Origin of Eukaryotic Cells (Benchmark Papers in Systematic and Evolutionary Biology, 9) (1986), Tracing the History of Eukaryotic Cells (1994 – with Robert A. Obar), Explore the World Using Protozoa (1997 – coauthor), A Field Guide to Bacteria (2003) and Perl for Exploring DNA (2007 – with Mark D. LeBlanc).
I recently read about her book “A Field Guide to Bacteria” and wanted to know more about her view on “Bacteriocentrism,” and how one becomes “bacteriocentric.” I was so lucky to be introduced to Dr. Dyer, who was kind enough to accept to be interviewed for Micro Writers, and immediately answered my questions about her book and her student project, Genomics Research Group, via email. So, here I am, sharing with you this very interesting interview.
1. Why did you decide to write the book “A Field Guide to Bacteria”? Field microbiology and symbiosis are among your research interests ; is this one of the reasons that made you dig deeper in bacterial populations and “think like a microbe”?
I first got the idea of a field guide when I was a graduate student and was very fortunate to be on a field microbiology expedition (in Baja California Mexico) with some world famous field microbiologists. I realized at once that a field guide should be written and one of them should write it and probably would! It did not occur to me that I would write it. It took me years to get to the point of being secure enough with my career. First I had to get a PhD, then a job, and then tenure. I also got married and had two children. Finally, about 15 years after that original idea, I realized that I had been accumulating enough information that I should begin to write. And so I did. However, I am still a bit surprised that nobody else wrote it.
I am naturally drawn to tiny things. I got a microscope for a present when I was 11 years old and it transformed some of my views of biology. I found that I loved the microscopic world. But I also like miniatures in general such as tiny furniture and dishes and things in dolls houses. I have in my library at home, some shelves devoted to a doll house and two miniature rooms.
2. What is meant by “becoming bacteriocentric”? And how does this lead to better understanding of the biology of bacteria?
We humans are mostly visual and auditory, the primary senses by which we perceive and analyze the world. It is probably impossible for us to be otherwise. Furthermore, we are gigantic and multicellular and terrestrial in marked contrast to the vast majority of organsims on Earth. Nonetheless, I think it is an excellent exercise for any biologist at least to try bacteriocentricity. The bacterial or microbial world is primarily olfactory and tactile. They are single celled (intimate with their environments), tiny and aquatic. I cannot avoid being anthropocentric but I can at least be more aware of the limitations of my size, habitat, and senses. My goal is to have as much humility as I can manage when I observe the world of microbes.
3. “Many groups of bacteria can be easily identified in the field (or in the refrigerator) without a microscope” and “Bacteria can be seen and smelled”, as a pharmacy student, I want ask how could that be achieved?
Well, do you have the book yet? There are many examples but the basis of all of them is that bacteria, when they are in an appropriate environment, are likely to do quite well: reproducing abundantly, taking in and transforming molecules, sending out wastes. In many cases (surprisingly many) the abundance is on a level perceptible by humans. The field marks just need to be revealed and interpreted. Otherwise, they may be easily overlooked or misunderstood. My first experience with this as a graduate student was being shown the distinctive pigmentations and odors of bacteria in sulfur cycles in Baja California.
4. You have a project with Dr. Mark LeBlanc, professor of Computer Science, called “Wheaton College Genomics Group.” Why did you do such a project for undergraduate students? How did it raise their potential?
One day about ten years ago, Mark asked me if I had any large datasets that his computer science students might analyze in their course on algorithms. It happens that I teach genetics and am fascinated with genomes. At that time, genome sequences were becoming more available at NCBI. I had not realized that it would be so easy to collaborate with a computer scientist. I had lots of questions about genomes and we just started right in with devising some answers. Right now, we are interested in characterizing horizontal transfer events of distantly related bacteria and archaea. There are hundreds of complete microbial genomes at NCBI and most have not been completely analyzed. Therefore, there is plenty for us and our students to do.
We ended up writing a book on the topic because we wee in need of a text that could be used both by biologists and computer scientists.
Tags: A Field Guide to Bacteria, bacteriocentric, bacteriocentrism, Betsey D. Dyer, biologists, computer scientists, field microbiology, Mark LeBlanc, NCBI, Perl, Perl for Exploring DNA, sulfur cycles, symbiosis, Wheaton College Genomics Group
 1 Comment »
1 Comment »
Yediot Ahronot (Literally: Latest News)
Cure for radiation sickness found?
Published: 07.17.09
A team of scientists has succeeded in developing an anti-radiation. “The process that led up to the medical innovation dates back to 2003, when Professor Gudkov –the head of the team- came up with the idea of using protein produced in bacteria found in the intestine to protect cells from radiation.” Mice received that purified protein survived the amount of radiation that killed the control group.
What kind of news is that?! Do that legendary bacteria and that miraculous protein actually have names?! If so, why don’t newspapers include it? Should bloggers do everything?!
Professor Andrei Gudkov – Chief Scientific Officer at Cleveland BioLabs, is interested in protecting cells against apoptosis induced by cancer therapy as well as radiotherapy. He worked on p53 –the famous tumor suppressor- and found that p53 has a role in inducing apoptosis. The research group suggested that p53 inhibitors can protect normal cells against chemo- and radiotherapy, and it’s been found that it sensitizes tumor cells to the therapy (PMID: 15865929). They also showed that PFTmu (pifithrin-mu), a small isolated inhibitor of p53, protected primary mouse thymocytes from p53-induced apoptosis caused by radiation (PMID: 18403709).
The breakthrough discovery mentioned above has been published in Science – 11 April 2008. It is about the injection of “flagellin” purified from Salmonella enterica serovar Dublin into mice and monkeys. It causes suppression of apoptosis by binding to Toll-like receptor 5 (TLR5) and activation of the nuclear factor–kappaB (NF-kappaB) pathway, the same mechanism used by tumor cells to inhibit the function of the p53 pathway (PMID: 18403709). To reduce its immunogenecity and toxicity, they engineered a polypeptide derived from flagellin with the “important” domains only, N and C termini separated by a linker. The engineered protein (named: CBLB502) was found to provide radioprotection in rhesus monkeys and mice against lethal doses of gamma-radiation and accompanied hematopoietic system and gastrointestinal tract acute radiation syndrome, with no alteration of the efficacy of the radiotherapy.
Tags: Andrei Gudkov, anti-radiation, flagellin, NF-{kappa}B, p53 inhibitor, PFTmu, pifithrin, radioprotective, radiotherapy, Salmonella enterica, TLR5, Yediot Ahronot
 No Comments »
No Comments »
When you hear/ read the term “Phage Therapy“, you’ll be automatically directed to the concept of using bacteriophages, the virus-like particles that infect bacteria, to kill/ lyse the resistant bacterial strains, instead of the “useless” antibiotics that allowed bacteria to fool them & develop resistance against them. The initial target of phage therapy was to kill the bacteria using phages; because they act like any other virus; get in, multiply and lyse the cell. But, by this way, bacteria develop resistance against phages more rapidly. So, they may become useless by time. In this paper from PNAS: “Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy,” two bioengineers, Timothy K. Lua and James J. Collins, from Boston University successfully engineered the Enterobacteria filamentous phage M13 to weaken bacteria not to kill it. Sounds strange, right? By engineering M13, they gave us a variety of options:
1st, we may make M13 overexpress a bacterial protein named lexA3 which inhibits the ability of the bacteria to repair their damaged DNA by the action of Ofloxacin –as pharmacophils, who had 2 consecutive chemotherapeutics courses, we may recall that quinolones’ MOA is generation of ROS. So, the repressor suppresses the bacterial SOS mechanism. Very promising results were observed; the adjuvant therapy increased the survival rate of mice infected with resistant E. coli. It was also observed that the adjuvant therapy reduced the rate of developing mutations/ resistance within the E. coli population.

2nd, bacteriophage can be responsible for expression of certain proteins that can attack gene networks in bacteria which are not target for existing antibiotic classes. I will mention just one example here, expression of CsrA which is a “global regulator of glycogen synthesis and catabolism, gluconeogenesis, and glycolysis, and it also represses biofilm formation,” biofilms is thought to be related to antibiotic-resistance and OmpF porin which is used by quinolones to enter the bacterial cell, it may enhance its entrance.

Now, thanks to the engineered phages, we can use the old beloved antibiotic classes to treat bacterial infection using the engineered phages as an adjuvent therapy to potentiate the cidal action of the antibiotic on the former-resistant strains. A precaution was made to ensure that no lysogeny would take place in the human cells is that the phages were engineered to be “nonreplicative”. But we still have two problems regarding Phage Therapy in general: identifying the strain responsible for the infection & making sure that the human immune system won’t elicit an immune response against phages, they’re “foreigners” after all!
Image credits:
1- “Schematic of combination therapy with engineered phage and antibiotics. Bactericidal antibiotics induce DNA damage via hydroxyl radicals, leading to induction of the SOS response. SOS induction results in DNA repair and can lead to survival. Engineered phage carrying the lexA3 gene (lexA3) under the control of the synthetic promoter PLtetO and an RBS acts as an antibiotic adjuvant by suppressing the SOS response and increasing cell death”: http://www.pnas.org/content/106/12/4629.figures-only
2- “CsrA suppresses the biofilm state in which bacterial cells tend to be more resistant to antibiotics. OmpF is a porin used by quinolones to enter bacterial cells. Engineered phage producing both CsrA and OmpF simultaneously (csrA-ompF) enhances antibiotic penetration via OmpF and represses biofilm formation and antibiotic tolerance via CsrA to produce an improved dual-targeting adjuvant for ofloxacin”: http://www.pnas.org/content/106/12/4629.figures-only
Tags: antibiotic, bacteriophage, bioengineering, engineered bacteriophage, M13, ofloxacin, phage, phage therapy, resistant
 2 Comments »
2 Comments »
Streptococcus pneumoniae is a pathogen normally found in nasopharynx of about 5-10 % of healthy people. Despite its name, it can cause many other diseases as meningitis, septic meningitis, otitis media, & osteomyelitis.
S. pneumoniae main virulence factors are:
a) Polysaccharide capsule: that inhibits phagocytosis.
b) Pneumolysin: a protein that causes lysis of host cells & activate complement.
c) Hydrogen peroxide: causes damage to host cells.
d) Pili: contribute in colonization of upper respiratory tract and increase the formation of large amounts of TNF.

Researchers led by Jesús M. Sanz at the Miguel Hernandez University and Maarten Merkx at the Eindhoven University of Technology have now introduced a highly promising new approach for the development of drugs to treat pneumococci.
The cell wall of pneumococci contains special polymers, called teichoic acids that are rich in choline which act as a docking station for a number of special proteins that are involved in important processes as:
1) Cell wall division.
2) Release of bacterial toxins.
3) Adhesion to infected tissues.
If choline is added to a culture of pneumococci, the molecules occupy the choline binding sites of the CBPs so that the proteins can no longer bind to the cell walls of the pneumococci.
The bacteria continue to multiply but they can’t separate from each other, these results in long chains of linked cells. Also, the toxin released is stopped.
However, choline is not suitable to be used as a drug because an effective dose would be far too high.
The researchers thus developed the foundation for a new drug that binds CBP much more strongly than individual choline molecules. The drug imitates the choline architecture of the cell wall by presenting multiple choline groups. This will occupy multiple choline binding sites of the CBP.
Also, the required dosage of this CBP inhibitor lies within the range that is acceptable for pharmaceuticals.
Source: Science Daily.
Image credits: S.pneumoniae.
Image credits: S.pneumoniae.
Tags: CBPs, choline binding sites, meningitis, osteomyelitis, otitis media, pili, Pneumolysin, Streptococcus pneumoniae, teichoic acids, TNF
 1 Comment »
1 Comment »
Hello, hello. You’re now tuned to your favorite blog: micro-writers.egybio.net. Tonight we have this very special guest, live, online. After two months of waiting, we finally got this exclusive interview with the emerged Streptococcus pyogenes strain, the most dangerous ever, M1T1. We have it here, with us, in the studio.
– Hello, M1T1. Welcome in our studio.
– Hey there.
– We knew from our resources, which are totally classified, that you got yourself in trouble recently.
– (Interrupting), I did NOT get myself in trouble. EID set me up.
– M1T1, Would you please calm down & tell us a little more about yourself?
– Well, I belong to Group A streptococci (GAS) aka Streptococcus pyogenes. M1T1 is my serotype; I’m just a clonal strain. As you know, S. pyogenes colonize human skin & throat causing either non-invasive (sore throat, tonsillitis & impetigo) or invasive (necrotizing fasciitis NF, scarlet fever & streptococcal toxic-shock syndrome STSS) infections. Actually, NF gave me my nick: Flesh-eating bacteria.
– So, you cause all people NF & STSS?
– No, kid. It depends on their genetic susceptibility, what you call “Host–pathogen interactions”. I was isolated from patients with invasive as well as non-invasive infections during 1992–2002. This is NOT entirely my fault; humans can make me extra virulent by selecting the most virulent members.
– Back to your history, when have you exactly been isolated?
– M1 & her sisters were the worst nightmare in US & UK in the 19th century as they caused the famous pandemic of scarlet fever. “Nevertheless”, early 1980s was the golden age of my strain as well as my very close sisters M3T3 & M18. We caused STSS & NF in different parts of the world. Great times, great times!
– Only for you, I suppose! So, what made you hypervirulent? What caused you this “epidemiologic shift”?
– Two reasons Dr Ramy K. Aziz identified that improved my fitness to humans: the new genes I got from phages & “host-imposed pressure”. Both resulted in the selection & survival of me M1T1 the hypervirulent strain. Dr Aziz’s work at Dr Kotb’s lab resulted in identification of a group of genes I got from phages that changed my entire life.
– Interesting! Tell us more about that. How did phages “change your life”?
– Dr Aziz proved that I differ from my ancestral M1 when he found that I have 2 extra prophages (lysogenized phages didn’t get the chance to lyse me, so they became integrated in my genome):
1. SPhinX which carries a gene encodes the potent superantigen SpeA or pyrogenic exotoxin A (scarlet fever toxin).
2. PhiRamid which carries another gene encodes the most potent streptococcal nuclease ever, Sda1.
3. He also found that phages conversion from the lytic state to the lysogenic state resulted in exchange of toxins between our different strains (aka Horizontal Gene Transfer). Phages are very good genetic material transporters, what makes “strains belonging to the same serotype may have different virulence components carried by the same or highly similar phages & those belonging to different serotypes may have identical phage-encoded toxins.” What a quote from Rise and Persistence of Global M1T1 Clone of Streptococcus pyogenes.
– Well, It was not that interesting. So, what? What’s the significance? How that made you hypervirulent?
– You can’t get it? You’re not that smart, are you? Tell me, what made M1 hypervirulent causing scarlet fever in the 1920s and me hypervirulent causing STSS in the 1980s with a 50-years decline period?
– Superantigen?
– Exactly. You do have your moments! Superantigen encoding-gene was present in us and absent in strains isolated in the period between them. The interesting part, for me of course, that humans after 50 years of absence of hypervirulent strains had absolutely no superantigen-neutralizing antibodies. That was the real invasive party. Superantigen causes high inflammatory response because of its non-specific binding to immune system components (antibodies & complements) causing an extremely high inflammatory response. In fact, SuperAg inflammatory response is “host-controlled”.
– So, what about Sda1?
– Streptodornase (streptococcal extracellular nuclease) helps me to degrade neutrophils that entrap me in the neutrophil extracellular traps (NETs). So, I can invade humans freely & efficiently and be able to live in their neutrophils. Dr Aziz proved in his paper “Post-proteomic identification of a novel phage-encoded streptodornase, Sda1, in invasive M1T1 Streptococcus pyogenes” that it’s all about C-terminus in my Sda1; the frame-shift mutation increased my virulence while deletion decreased it.
– Now we know about your SuperAg & nuclease (DNase), what’s the “host-imposed pressure”?
– I have my own SpeB (Protease), I use it to degrade my other proteins (virulence factors), which provides me with a good camouflage & gives me access to blood. When the host immune system recognizes me, it traps me in NETs. At this time, I secret Sda1 to degrade neutrophils. Actually, SpeB protects you, humans, from my Sda1& my other toxins. When SpeB was compared in patients with severe & non-severe strep infections, it was found that SpeB wasn’t expressed in case of severe infection. Expression of SpeB may be host-controlled, as host selects the mutants with a mutation in covS, a part of my regulatory system which regulates my gene expression including SpeB gene.
– Finally, M1T1. How do you see your future?
– More new phage-encoded genes, more selection of the hypervirulent strains by the host & more regulation of expression of my virulence factors. Pretty good future! I also count on humans to not develop immunity against me like what happened in 1980, when I got new virulence factors or allelic variations in my old ones.
Thank you, M1T1. Pleasure talking to you…….M1T1? M1T1, where are you? Why do I feel this strange pain in my throat?
Image credits:
Streptococcus pyogenes: http://adoptamicrobe.blogspot.com/
Tags: covS, epidemiologic shift, flesh-eating bacteria, global M1T1 clone, group A streptococci (GAS), horizontal gene transfer, host-imposed pressure, host–pathogen interactions, impetigo, M1T1, Malak Kotb, necrotizing fasciitis, neutrophil extracellular traps (NETs), phage-encoded toxins, pyrogenic exotoxin A, Ramy K. Aziz, S. pyogenes, scarlet fever, sore throat, streptococcal nuclease Sda1, streptodornase, STSS, superantigen SpeA, tonsillitis
 5 Comments »
5 Comments »
Who could possibly believe that our anaerobic spore-forming Clostridia will be used as an anti-tumor therapy?! It’s called Clostridium-based tumor targeted therapy. “Give me a break,” that’s exactly what I said when I read the review of the new book Clostridia: Molecular Biology in the Post-genomic Era.
So, how does it work? There are various non-pathogenic Clostridia strains which could replicate within solid tumors upon systemic administration. The interesting part is coming right up: Why solid tumors?! It’s because of its very unique physiology; it characterized by hypoxia & necrosis which totally fits the anaerobic Clostridia. The advantage will be the selectivity & targeting of the cancer cells leading to destroying them.
It was news to me to know that Cl. perfringens causes food poisoning like any food-borne illness & causes antibiotic-associated diarrhea (When I hear the name Cl. perfringens, I orient myself toward gas gangrene right away). Its enterotoxin gene (cpe) is present on the chromosome itself (in food poisoning isolates) and on the plasmid (in the antibiotic-associated diarrhea isolates). The enterotoxin binds to claudin receptors, then there’s oligomerization or “prepore” formation & finally prepore insertion takes place to form the functional pore which kills the cells by apoptosis. So CPE/CPE derivatives could be used for cancer therapy.
Tags: anti-tumor, clostridia, Clostridium-based tumor targeted therapy, CPE/CPE, enterotoxin, solid tumors
 2 Comments »
2 Comments »
 Researchers at Yale & the University of Chicago were faced with a surprising conclusion based on their scientific experimentation on mice. Unlike the common belief that microbes are in fact “bad” & possess a harmful threat to our health, some of these bacteria prove their innocence. Mice, that were exposed to common bacteria in the normal gut flora, were protected against the development of Type I diabetes. Previous research had shown that mice, exposed to killed Mycobacterium tuberculosis, were also protected. So, this means that mice that grow in their natural habitat are better off than the ones raised in the much improved sanitary conditions of the lab. Researchers at Yale & the University of Chicago were faced with a surprising conclusion based on their scientific experimentation on mice. Unlike the common belief that microbes are in fact “bad” & possess a harmful threat to our health, some of these bacteria prove their innocence. Mice, that were exposed to common bacteria in the normal gut flora, were protected against the development of Type I diabetes. Previous research had shown that mice, exposed to killed Mycobacterium tuberculosis, were also protected. So, this means that mice that grow in their natural habitat are better off than the ones raised in the much improved sanitary conditions of the lab.
This comes to support the hypothesis many scientists have lately adopted. They tend to believe in a directly proportional realtionship between a person’s exposure to parasites, bacteria, worms, etc.. within the surrounding evironment and his immunity. The more, the better..that is within limits of course.
This actually makes perfect sense to me. It is really obvious when you see, for example, people living in third world countries with mosquitoes hovering around and considered normal. But when they travel abroad for a while and come back, they get different sorts of allergies & rashes from those previously “harmless” mosquitoes. What parasites and microbes do for you is not all bad. Unfortunately, I had to experience this dilemma.
Source: ScienceDaily
Tags: bacteria, diabetes, flora, immunity, mice, tuberculosis, Yale
 No Comments »
No Comments »
Sept 16th:
Hi, Diary;
Now it’s in the press. It’s humiliating. In Scientific American, it was written under the title: Turning Bacteria into Plastic Factories. So that’s what we, E. coli, turned into: slaves for humans to do whatever they want us to do.
I remember this like it happened yesterday. It was someday in the beginning of the year 2008. That day we found ourselves in those tanks, large ones, filled with sugar & water. “This is great; we can ferment the whole amount of sugar in the tank. This is the perfect place to live in,” that is what we thought. Then we got this strange order from our genomes or plasmids, I’m not sure & found ourselves producing that enormous amount of 1, 4- Butanediol (BDO). This was so strange because BDO used to be toxic to us at low levels (I’ve heard once that any production of a non-native material inhibits our growth). Then we realized the truth; we’ve been genetically engineered to tolerate BDO, the raw material for a very large number of plastic, rubber and fiber products including solvents, fine chemicals, pharmaceuticals, automotive components, electrical and electronics components, as well as apparel fibers.
Honestly, I don’t know. Sometimes I feel that bioengineers from San Diego-based Genomatica, Inc. are right after all; they modified us, they wanted to make use of us, they wanted to save money & energy needed to produce BDO from non-renewable petrochemical feedstocks, the currently used method. Plus it causes us no harm as Bioengineer Christophe Schilling, president and co-founder of the company said: “We have engineered the organism such that it has to secrete that product in order for it to grow.”
Now I know what it feels like with my fellow bacterial species, the natural born producers. Most bacteria synthesize the organic polyesters Polyhydroxyalkanoates (PHAs) to be used as a carbon & energy storage material. Now they’re discussing the ability to use these PHAs as biodegradable plastics.
 Source: A piece of a plastic notebook found in the Petri dish appears in the picture below, Genomatica labs, San Diego. Source: A piece of a plastic notebook found in the Petri dish appears in the picture below, Genomatica labs, San Diego.
Image credits:
Genetically engineered E. coli may produce plastics: http://www.sciam.com/
Tags: BDO, biodegradable plastics, Christophe Schilling, genetically engineered, Genomatica, PHAs
 3 Comments »
3 Comments »
 Recently, I have been able to get in touch with Professor Jan Roelof van der Meer after reading about his work in EurekAlert in the field of color-coded bacteria. Currently, he is an associate professor at the Department of Fundamental Microbiology, University of Lausanne. Recently, I have been able to get in touch with Professor Jan Roelof van der Meer after reading about his work in EurekAlert in the field of color-coded bacteria. Currently, he is an associate professor at the Department of Fundamental Microbiology, University of Lausanne.
In a review published in collaboration with Professor Robin Tecon, the researchers explained why the bacteria were specifically useful in the detection & tracing back the age of oil spills, chemicals and other pollutants leaking into seawater and the soil. Since the bacteria are easily manipulated, researchers were able to genetically produce MBS “Microbe-Based Sensors” which produce specific reporter proteins when in contact with a certain pollutant. Such reporter proteins can then be detected merely by observation or instrumentally.
Enclosed within the review, this figure illustrates the concept of a bacterial sensor-reporter cell where the benzene-ring-look-a-likes represent the pollutants.
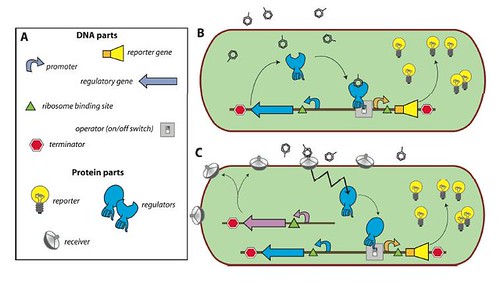
And I leave you with the interview:
1. Is there hope that MBS won’t just play a role in detection, but in cleaning up as well?
Normally not. To enhance biodegradation rates in the environment, one usually tries to stimulate the bacteria which are already present at the site. There are no cases where genetically modified bacteria were applied to clean up contamination.
2. How can this method trace back the age of a spill?
Interestingly, we found that there seems to be a specific pattern of dissolution of different compounds from oil. A fresh spill will first ‘show’ linear alkanes and compounds like benzene, toluene, ethylbenzene. Only later will polycyclic aromatic hydrocarbons, like naphthalene, appear. We had not seen this before, because one typically cannot measure the first phase of an oil spill, since this is detected only after a while.
3. Which method do you prefer in the genetic engineering of these MBS?
Depends. For E. coli, we use very classical cloning techniques involving plasmids. For other bacteria, we have to use transposon delivery methods mostly.
4. Is the acquired trait of producing a reporter protein passed on to future generations of the bacteria?
Normally yes. If the reporter construct is integrated in the genome of the bacteria, it is relatively stably maintained even without selection pressure for the marker. When the construct is on a plasmid, like in E. coli, one has to constantly keep the ‘pressure’ for the marker on the plasmid, usually an antibiotic resistance marker.
5. This field, as you kindly mentioned, started 20 years ago; what hope lies for its progess in the future?
My major hope is that people (industries, labs) finally apply the methods in their analysis as alternatives for costly chemical analysis. Further progress has to come from miniaturization, multiple target detections and improved methods to preserve the bacterial cells.
Tags: bacteria, biosensor, color coding, genetic engineering, Lausanne, MBS, oil spill, reporter protein, van der Meer
 2 Comments »
2 Comments »
“Did you brush your teeth today ??”
I think you will ask this question to yourself & your family daily.
When you know that bad oral hygiene will drive you to heart disease, many of us may change his opinion.
Simply, people who don’t brush their teeth regularly will start with inflammation of their gums “Gingivitis“.
If this situation continues, a chronic inflammation occurs “Periodontal Disease“ with a bleeding gum.
Bleeding gum is a great risk factor, as it provides an entry to the blood stream for 700 different types of bacteria.
Streptococcus gordonii and Streptococcus sanguinis are two common infecting agents.
After they reach the blood stream, they bind with blood platelets, this creates two problems:
First: If we use an oral antibiotic to kill oral bacteria thus preventing them from reaching the blood stream, by time bacteria become resistant.
Second: The binding of bacteria with platelets encases them & shields bacteria from our immune system till they reach the heart causing heart diseases.
So, if we prevent the binding between bacteria & platelets, this will help our immune syst em kill bacteria & protect our heart. em kill bacteria & protect our heart.
“We are currently in the process of identifying the exact site at which the bacteria stick to the platelets” said Professor Howard Jenkinson. “Once this is identified we will design a new drug to prevent this interaction.”
Source: Bristol University news.
Image credits: Periodontal Disease
Image credits: Professor Howard Jenkinson
Tags: bleeding gum, Gingivitis, Heart Attack, Periodontal Disease, Streptococcus gordonii, Streptococcus sanguinis
 3 Comments »
3 Comments »
|
 from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including
from Boston University. Among her research interests are symbiosis, evolution of cells, field microbiology and genomics. She is also part of the Genomics Research Group, a student project that she launched in collaboration with Mark LeBlanc, professor of Computer Science. Dr. Dyer has written several books, including 










 em kill bacteria & protect our heart.
em kill bacteria & protect our heart.

 Entries (RSS)
Entries (RSS)