Interview: Professor Jan van der Meer
Posted by: Radwa Sharaf in Bacteriology, Interview, Meet a Scientist Recently, I have been able to get in touch with Professor Jan Roelof van der Meer after reading about his work in EurekAlert in the field of color-coded bacteria. Currently, he is an associate professor at the Department of Fundamental Microbiology, University of Lausanne.
Recently, I have been able to get in touch with Professor Jan Roelof van der Meer after reading about his work in EurekAlert in the field of color-coded bacteria. Currently, he is an associate professor at the Department of Fundamental Microbiology, University of Lausanne.
In a review published in collaboration with Professor Robin Tecon, the researchers explained why the bacteria were specifically useful in the detection & tracing back the age of oil spills, chemicals and other pollutants leaking into seawater and the soil. Since the bacteria are easily manipulated, researchers were able to genetically produce MBS “Microbe-Based Sensors” which produce specific reporter proteins when in contact with a certain pollutant. Such reporter proteins can then be detected merely by observation or instrumentally.
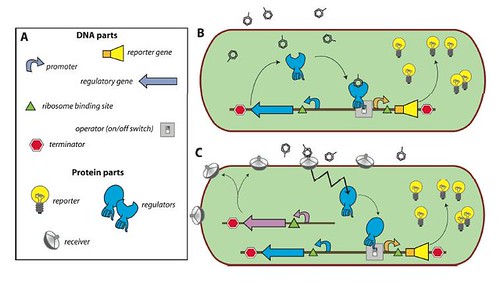
Enclosed within the review, this figure illustrates the concept of a bacterial sensor-reporter cell where the benzene-ring-look-a-likes represent the pollutants.
And I leave you with the interview:
1. Is there hope that MBS won’t just play a role in detection, but in cleaning up as well?
Normally not. To enhance biodegradation rates in the environment, one usually tries to stimulate the bacteria which are already present at the site. There are no cases where genetically modified bacteria were applied to clean up contamination.
2. How can this method trace back the age of a spill?
Interestingly, we found that there seems to be a specific pattern of dissolution of different compounds from oil. A fresh spill will first ‘show’ linear alkanes and compounds like benzene, toluene, ethylbenzene. Only later will polycyclic aromatic hydrocarbons, like naphthalene, appear. We had not seen this before, because one typically cannot measure the first phase of an oil spill, since this is detected only after a while.
3. Which method do you prefer in the genetic engineering of these MBS?
Depends. For E. coli, we use very classical cloning techniques involving plasmids. For other bacteria, we have to use transposon delivery methods mostly.
4. Is the acquired trait of producing a reporter protein passed on to future generations of the bacteria?
Normally yes. If the reporter construct is integrated in the genome of the bacteria, it is relatively stably maintained even without selection pressure for the marker. When the construct is on a plasmid, like in E. coli, one has to constantly keep the ‘pressure’ for the marker on the plasmid, usually an antibiotic resistance marker.
5. This field, as you kindly mentioned, started 20 years ago; what hope lies for its progess in the future?
My major hope is that people (industries, labs) finally apply the methods in their analysis as alternatives for costly chemical analysis. Further progress has to come from miniaturization, multiple target detections and improved methods to preserve the bacterial cells.



 Entries (RSS)
Entries (RSS)